pyRinocitologia
Central repository for Rinocitologia project developed as an undergraduate thesis. The purpose of this repository is to contains all development code allowing...
Design Philosophy
- Reproducibility
- Process automation
- Explicit is better than implicit
- Infrastructure built around data
- Prefer cross-platform solution
Directory Structure
.
|
+--- data data used by code
|
+--- models pre-trained models and stored weights
|
+--- notebooks jupyter notebooks for analisys and experiments
|
+--- report resources used for documentation
|
+--- src main code
|
+--- tools code for misc utilities
|
+--- config.ini.example configuration file
|
+--- README.md
|
+--- requirements.txt python pip dependencies file
Setup
-
Create a virtual environment with conda or virtualenv (optional but recommended)
-
Install dependencies:
pip install -r requirements.txt
- Setup the environment with config.ini, rename config.ini.example into config.ini and change content to adapt to your environment
Use
You can use your custom data or use our Open Data Dataset. dataset/s could be clone directly in the main repository.
Data Gathering
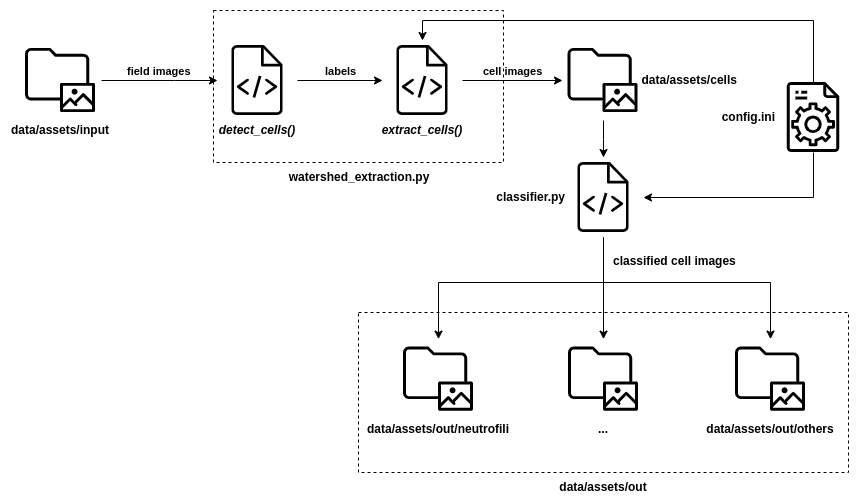
Execution Flow
Tools
Various utilities script has been developed.
- metadata_generator.py is a small command line utility to help in generate a dataset metadata json file. for more info:
python metadata_generator.py --helpnote:
image annotation tool : VGG Image Annotation (VIA)
Warnings
The python interpreter cwd (Current Working Directory) must be set to the root directory/parent directory of /src folder eg:
import os
os.chdir("..")or equivalently, call python interpreter from the parent directory:
python <parent dir>/src <script name>or add to PYTHONPATH the parent directory:
export PYTHONPATH=<root directory>
python <script name>(if you use pyCharm IDE this is done automatically in run configuration)