Currently, this code is dependent on experimental features of the TensorKit project. So you need to install the block_optimizer branch. I aim to merge this into master soon to make the installation process easier. For now, run the following two lines to install the correct versions of tensorkit and bcd_tensorkit.
pip install https://github.com/MarieRoald/TensorKit/archive/block_optimizer.zip
pip install git+https://github.com/MarieRoald/bcd_tensorkit.git
import tenkit
import bcd_tenkit
import numpy as np
import matplotlib.pyplot as pltCreate a decomposition for a set of coupled matrices with non-negative factors that follow the PARAFAC2 constraint
I, J, K = 20, 30, 40
rank = 3
A = np.random.standard_normal(size=(I, rank))
A = np.maximum(A, 0)
C = np.random.uniform(size=(K, rank))+ 0.1
blueprint_B = np.random.standard_normal(size=(J, rank))
blueprint_B = np.maximum(blueprint_B, 0)
B = np.empty(shape=(K, J, rank))
for k in range(K):
B[k] = np.roll(blueprint_B, shift=k, axis=0)
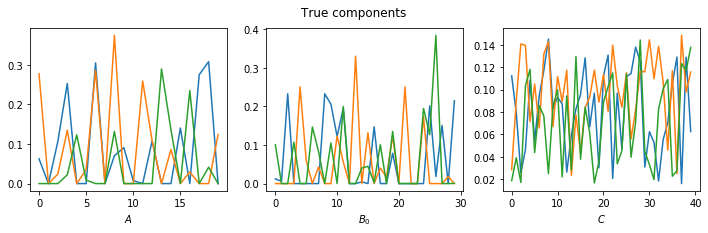
fig, axis = plt.subplots(1,3, figsize=(12,3))
axis[0].plot(A/np.linalg.norm(A))
axis[0].set_xlabel('$A$')
axis[1].plot(B[0]/np.linalg.norm(B[0]))
axis[1].set_xlabel('$B_0$')
axis[2].plot(C/np.linalg.norm(C))
axis[2].set_xlabel('$C$')
fig.suptitle('True components')
plt.show()X is a tensor with elements given by
decomposition = tenkit.decomposition.CoupledMatrices(A, B, C)
X = decomposition.construct_tensor() # Create tensor whose last mode represent matrix number
noisy_X = tenkit.utils.add_noise(X, 0.33)decomposer = bcd_tenkit.BCDCoupledMatrixDecomposer(
rank,
sub_problems=[
bcd_tenkit.Mode0ADMM(non_negativity=True),
bcd_tenkit.DoubleSplittingParafac2ADMM(non_negativity=True),
bcd_tenkit.Mode2ADMM(non_negativity=True)
],
max_its=1000,
convergence_tol=1e-8,
absolute_tol=1e-10,
print_frequency=100,
)decomposer.fit(noisy_X)
# Since noisy_X is a 3D numpy array, its matrices is extracted into a list
# like so: matrices = [noisy_X[:, :, k] for k in range(noisy_X.shape[2])] 100: The MSE is 0.310959, f is 4.10324, improvement is 5.81681e-08, coupling_errors: [2.797041014575114e-07, 3.7286601210992627e-05, 1.1582109416421031e-05, 0.0]
200: The MSE is 0.310959, f is 4.10323, improvement is 4.39264e-09, coupling_errors: [5.990262263456091e-08, 6.885537588476478e-06, 2.6606575848032075e-06, 0.0]
300: The MSE is 0.310959, f is 4.10323, improvement is 4.85884e-10, coupling_errors: [1.391739533501992e-08, 1.3417764302975869e-06, 6.254408375225476e-07, 0.0]
400: The MSE is 0.310959, f is 4.10323, improvement is 7.60386e-11, coupling_errors: [3.4006223509147945e-09, 2.7290285517803774e-07, 1.4968182529347784e-07, 0.0]
500: The MSE is 0.310959, f is 4.10323, improvement is 1.48148e-11, coupling_errors: [8.599414326123561e-10, 5.747019094758922e-08, 3.643730194742261e-08, 0.0]
600: The MSE is 0.310959, f is 4.10323, improvement is 3.2274e-12, coupling_errors: [2.2269642968672833e-10, 1.2467647259723803e-08, 9.017749253792294e-09, 0.0]
700: The MSE is 0.310959, f is 4.10323, improvement is 7.45266e-13, coupling_errors: [5.861378751430222e-11, 2.7773053600159416e-09, 2.2668882337945475e-09, 0.0]
800: The MSE is 0.310959, f is 4.10323, improvement is 1.78145e-13, coupling_errors: [1.5593954942078442e-11, 6.338686358207864e-10, 5.780112817822692e-10, 0.0]
900: The MSE is 0.310959, f is 4.10323, improvement is 4.39411e-14, coupling_errors: [4.177653827805391e-12, 1.479812357649442e-10, 1.4922954369970718e-10, 0.0]
estimated_decomposition = decomposer.decomposition
estimated_A = estimated_decomposition.A
estimated_B = estimated_decomposition.B # A list of factor matrices
estimated_C = estimated_decomposition.C
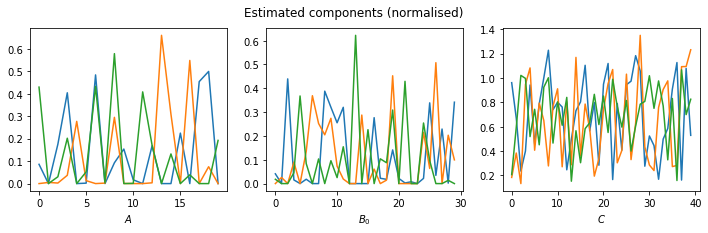
fig, axis = plt.subplots(1,3, figsize=(12,3))
axis[0].plot(estimated_A/np.linalg.norm(estimated_A, axis=0))
axis[0].set_xlabel('$A$')
axis[1].plot(estimated_B[0]/np.linalg.norm(estimated_B[0], axis=0))
axis[1].set_xlabel('$B_0$')
axis[2].plot(estimated_C/np.linalg.norm(estimated_C[0], axis=0))
axis[2].set_xlabel('$C$')
fig.suptitle('Estimated components (normalised)')
plt.show()Generating the components are done in the same way as the example code above.
import tenkit
import bcd_tenkit
import numpy as np
import matplotlib.pyplot as plt
I, J, K = 20, 30, 40
rank = 3
A = np.random.standard_normal(size=(I, rank))
A = np.maximum(A, 0)
C = np.random.uniform(size=(K, rank))+ 0.1
blueprint_B = np.random.standard_normal(size=(J, rank))
blueprint_B = np.maximum(blueprint_B, 0)
B = np.empty(shape=(K, J, rank))
for k in range(K):
B[k] = np.roll(blueprint_B, shift=k, axis=0)
decomposition = tenkit.decomposition.CoupledMatrices(A, B, C)
X = decomposition.construct_tensor() # Create tensor whose last mode represent matrix number
noisy_X = tenkit.utils.add_noise(X, 0.33)Fit a PARAFAC2 model using the AO-ADMM scheme
admm_logger = tenkit.decomposition.logging.CoupledMatricesFMSLogger(None, decomposition=decomposition)
admm_decomposer = bcd_tenkit.BCDCoupledMatrixDecomposer(
rank,
sub_problems=[
bcd_tenkit.Mode0ADMM(non_negativity=True),
bcd_tenkit.DoubleSplittingParafac2ADMM(non_negativity=True),
bcd_tenkit.Mode2ADMM(non_negativity=True)
],
max_its=1000,
convergence_tol=1e-8,
absolute_tol=1e-10,
loggers=[admm_logger]
)
admm_decomposer.fit(noisy_X)Fit a PARAFAC2 model using the flexible coupling scheme
flexible_logger = tenkit.decomposition.logging.CoupledMatricesFMSLogger(None, decomposition=decomposition)
flexible_decomposer = bcd_tenkit.BCDCoupledMatrixDecomposer(
rank,
sub_problems=[
bcd_tenkit.Mode0RLS(non_negativity=True),
bcd_tenkit.FlexibleCouplingParafac2(non_negativity=True),
bcd_tenkit.Mode2RLS(non_negativity=True)
],
max_its=1000,
convergence_tol=1e-8,
absolute_tol=1e-10,
loggers=[flexible_logger],
convergence_method='flex'
)
flexible_decomposer.fit(noisy_X)Plot the factor match score as a function of iteration number (stacking all B component matrices on top of each other beforehand)
plt.plot(flexible_logger.log_metrics, label="Flexible coupling")
plt.plot(admm_logger.log_metrics, label="AO-ADMM")
plt.legend()
plt.title("True component recovery")
plt.xlabel("Iteration")
plt.ylabel("FMS")
plt.xlim(0, 100)
plt.show()