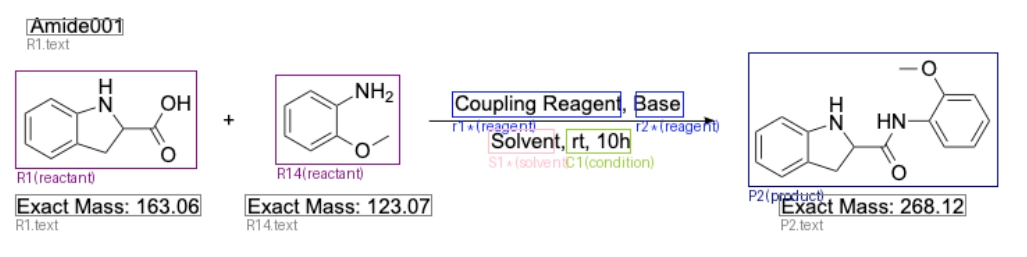
cdxml_tools is based on pure Python. Parse cdxml with graphical recognition to get the path of the reaction, condition, and the role of compounds.
A hypothetical usage scenario is to generate automated chemical experiment workflows or generate natural language descriptions. You can export cdxml and svg content from ChemDraw. Use the tools to get reaction info and recognition image.
The tool is based on pure Python. If you want the debug png feature, you need to install the following packages:
- Pillow
- wand
wand is a python binding from imagemagick. To install the package may need to install binary and set system env(see wand doc). The package is only used for converting svg to png.
The cdxml and svg content can export from ChemDraw Selection -> Get CDXML / Get SVG
- clone this repo
make install- see
tests/demo.ipynb
from cdxml import buildCdxml, parseCdxml
parseResult, img = parseCdxml(
cdxmlContent,
svg=svgContent,
withPosition=True, # parse result will have every node's coord
withCdxml=True, # parse result will have compound's cdxml fragment
withImg=True, # parse result will have compound's svg fragment
)The tools used the MIT license. Because the principle is a simple data converter. If you want to extend the feature or learn more about cdxml, highly recommend this article(CDXML format introduction).
Also welcome to submit issue or pull request :)
- add builder doc
- support configuration about graphical recognition rules