The paper is avaliable online in arxiv
This is the repo for paper 'Improving Contrastive Learning via Model Augmentation'. In this paper, we argue existing data augmentation methods for contrastive learning have the following weakness:
- optimal data augmentation methods are hard to devise,
- data augmentation methods destroy sequential correlations,
- data augmentation fails to incorporate comprehensive self-supervised signals.
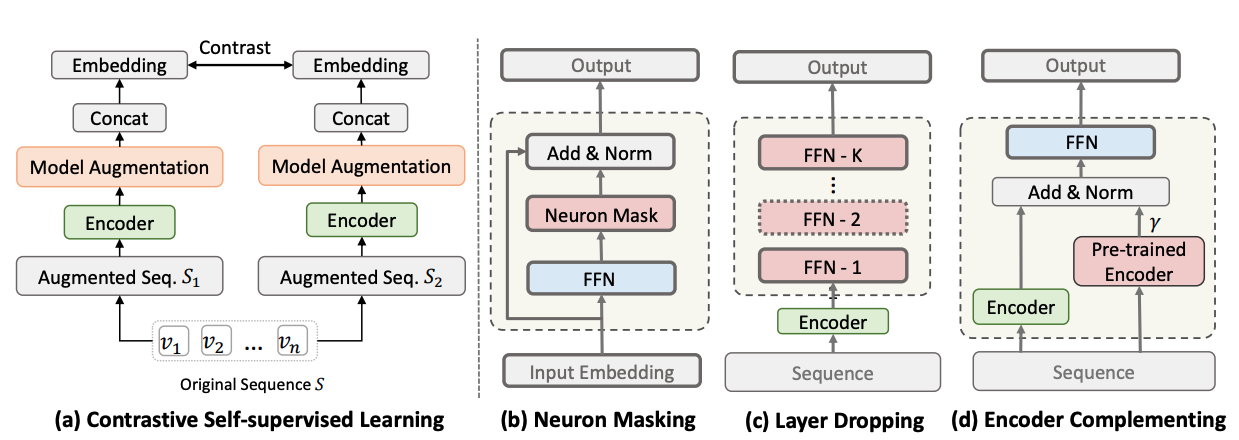
Therefore, we proposed three type of model augmentation methods:
neuron masking,layer droppingandencoder complementing.
If you use our code or follow our work, please cite as follows:
@article{liu2022srma,
title={Improving Contrastive Learning with Model Augmentation},
author={Liu, Zhiwei and Chen, Yongjun and Li, Jia and Luo, Man and Xiong, Caiming},
journal={arXiv preprint arXiv:2203.15508},
year={2022}
}
Python >= 3.7
Pytorch >= 1.2.0
tqdm == 4.26.0 faiss-gpu==1.7.1
We include the information of dataset in ./data/.
cd ./src/
- run with
python main.py --data_name Sports_and_Outdoors --gpu_id 0 --batch_size 256 --hidden_dropout_prob 0.5 --num_augmented_layers 2 --layer_drop_num 1 --model_aug_in_batch "same" --model_augmentationThis step will pretrain an encoder, which is saved in./output/. Note that the number indicating the--model_idxspecified inadd_argument. - Move the encoder to
./pretrain/and change the number to 'best'. We provide an example for Sports dataset. - run with
python main.py --data_name Sports_and_Outdoors --gpu_id 0 --batch_size 256 --model_augmentation --hidden_dropout_prob 0.5 --num_augmented_layers 2 --layer_drop_num 1 --use_pretrain --en_weight 0.5 --rec_model_aug
--data_name: the dataset to use, given in data--model_augmentation: whether to use model augmentation for sequential recommendation--hidden_dropout_prob: this is related to neuron masking, which is basically the masking probability in layers.--num_augmented_layers: this is related to layer dropping augmentation, which is implemented in the./src/modules.pyas in the classLayerDroplayer_drop: this is the other parameter in layer dropping augmentation, denoting how many augmented layers should be dropped during training. Should be a value less thannum_augmented_layers.use_pretrain: this is related toencoder_complementaugmentation, which adopts a pretrained encoder as a contrastive learning. More details should refer filetrainer.py, theSRMATrainer._cl_encoder()method.
- Current version of code doesn't support individual type of model augmentation. You may implement it by add other condition flags as in
main.py - We combine the data augmentation with model augmentation in this individual code. The data augmentation is based on our previous code CoSeRec
This model augmentation is related to the dropout ratio in encoder. Two arguments are related to Neuron Masking augmentation, which are hidden_dropout_prob and attention_probs_dropout_prob. The former controls the dropout probability in all hidden layers (such FC layer or embedding layer), while the later controls the dropout probability specifically for attention layers. Refer those self.dropout in file module.py
We implement the layer dropping mainly in the ./src/modules.py as in the class LayerDrop. The intuition behind layer dropping is to first append k additional layers after the encoder and then random drop m (args.layer_drop_num in the code) layers. We support append different types of layers, including single layer, intermediate layer, sasrec layer, etc. More details can be found in ./src/modules.py.
Besides the randomly m different layers, we also support contigent layer dropping. We first sample a value as the dropping probability for each layer, if it is greater than the given threshold, we will drop it. Otherwise, we will keep it. However, experiments doesn't support its effectiveness in our validation dataset. Welcome try in your own data and discuss with us!
We implement the encoder complementing by adding the contrastive loss between trained model and pre-trained model. Specifically, in code file trainers.py, the method function SRMATrainer._cl_encoder() specify this process:
# generate the embedding of the original sequence output from the pretrained encoder.
ori_sequence_output = encoder.transformer_encoder(original)
ori_sequence_flatten = ori_sequence_output.view(ori_sequence_output.shape[0], -1)
for slice in cl_output_slice:
cl_loss += en_weight*self.cf_criterion(slice, ori_sequence_flatten)encoder is from the pretrained encoder. original is the item sequence before data augmentation. We add the contrastive loss from two branches.
We reuse part of code from our previous work CoSeRec.