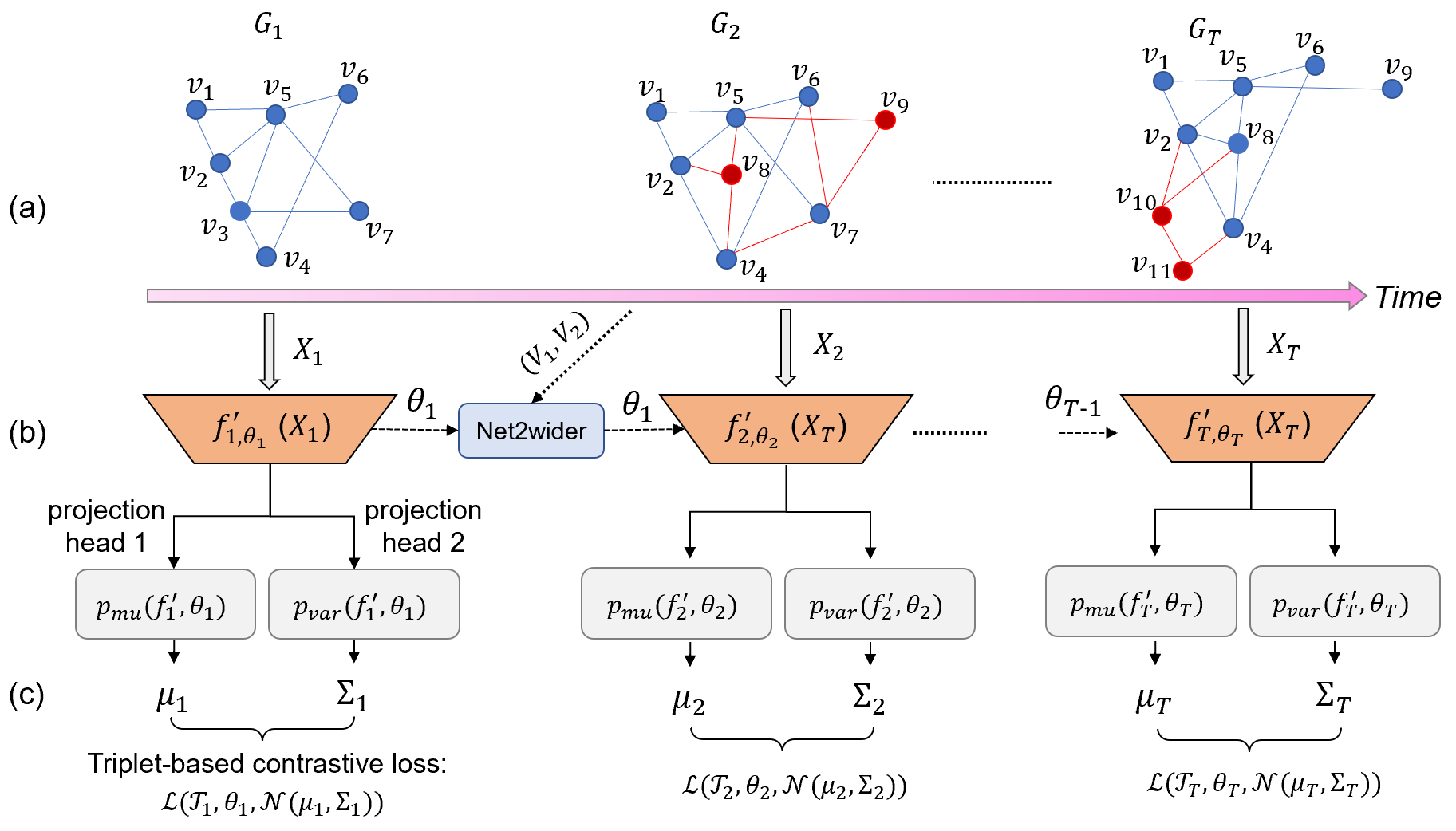
This repository contains the code for DynG2G: An Efficient Stochastic Graph Embedding Method for Temporal Graphs by Mengjia Xu, Apoorva Vikram Singh and George Em Karniadakis; This work has been submitted to IEEE Transactions on Neural Networks and Learning Systems (under revision). (The code was developed by Apoorva Vikram Singh and Mengjia Xu)
To create an environment to run the code, execute the shell file on the terminal using:
sh ./env_create.sh
The new environment called DynG2G is created. It will include all the dependencies needed to run the code. Activate it using:
source activate DynG2G
To download all datasets used in the experiment, run the shell file:
sh ./download_data.sh
To download the datasets seperately:
- AS dataset: Download link
- SBM dataset: Download link
- Bitcoin-OTC dataset: Download link
- UCI dataset: Download link
- Slashdot: Download link
- Facebook: Download link
- Reality Mining: Download link
- Digg: Download link
To generate the embeddings for AS dataset, run the command:
python Generate_Embeddings/AS_embed.py -f configs/[configFile]
Similarily, the embeddings for other datasets can be generated by running their corresponding files in Generate_Embeddings folder along with the desired configuration files (config).
- You can simply skip this process and download the embeddings from here.
To evaluate the embeddings for AS dataset run this command:
python Evaluation/eval-as.py -f configs/[configFile]
Similarily, we can evaluate the embeddings for other dataset by running their corresponding files in the Evaluation folder.
To plot the graphs for MAP and MRR, run the command:
python graph.py -f configs/[configFile] -d [datasetName]datasetName = as/sbm/otc/uci
This will generate the plot for MAP and MRR for the corresponding dataset.
If you find this code useful in your research, please cite:
@article{xu2021dyng2g,
title={DynG2G: An Efficient Stochastic Graph Embedding Method for Temporal Graphs},
author={Xu, Mengjia and Singh, Apoorva Vikram and Karniadakis, George Em},
journal={arXiv preprint arXiv:2109.13441},
year={2021}
url={https://arxiv.org/pdf/2109.13441.pdf}
}