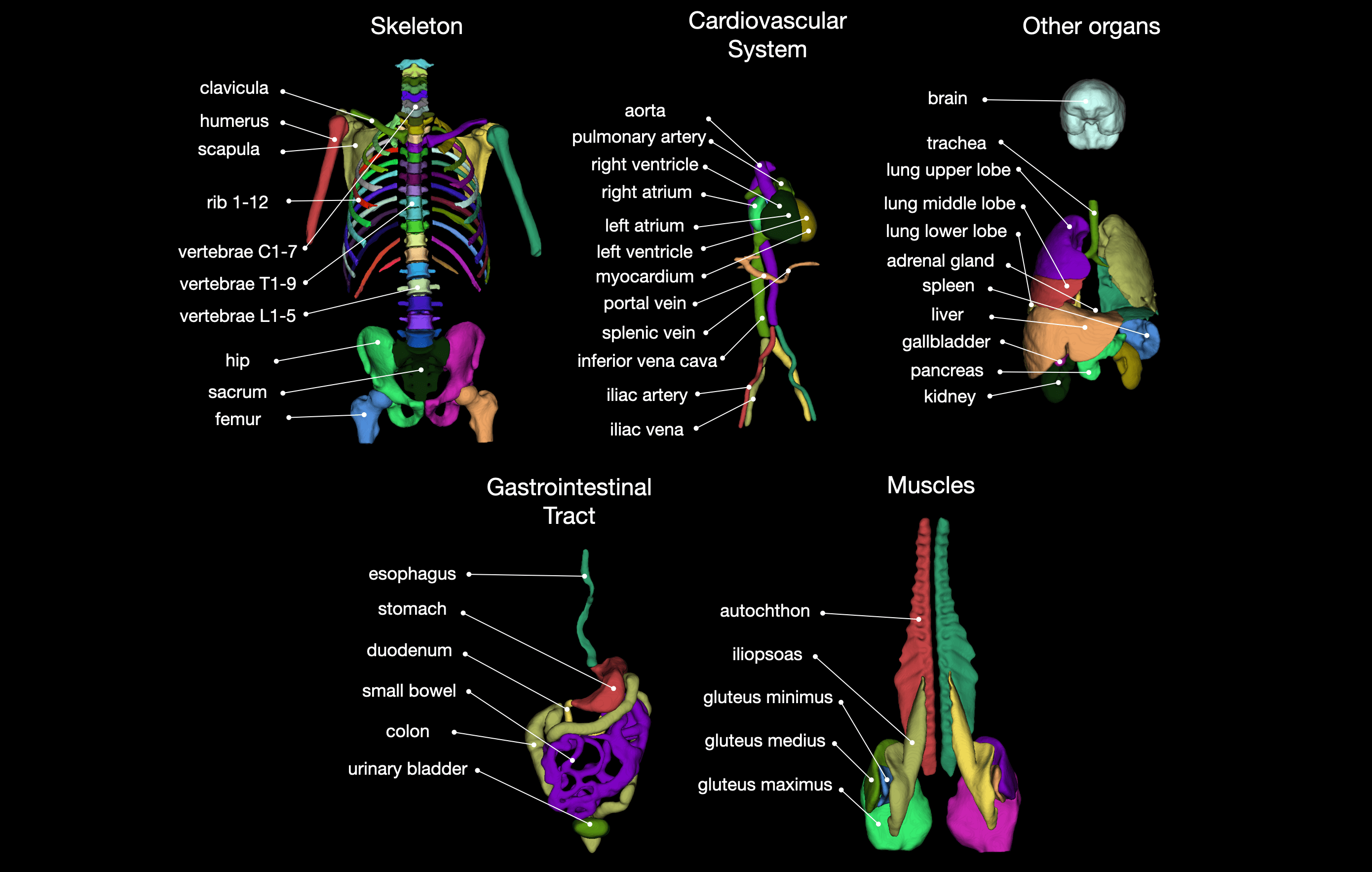
Tool for segmentation of 104 classes in CT images. It was trained on a wide range of different CT images (different scanners, institutions, protocols,...) and therefore should work well on most images. The training dataset with 1204 subjects can be downloaded from Zenodo. You can also try the tool online at totalsegmentator.com.
Created by the department of Research and Analysis at University Hospital Basel.
If you use it please cite our paper: https://arxiv.org/abs/2208.05868.
Install dependencies:
- Python >= 3.7
- Pytorch
- if you use the option
--previewyou have to install xvfb (apt-get install xvfb) - You should not have any nnU-Net installation in your python environment since TotalSegmentator will install its own custom installation.
- TotalSegmentator was developed for Linux or Mac. To make it work for windows see this comment.
Install Totalsegmentator
pip install TotalSegmentator
TotalSegmentator -i ct.nii.gz -o segmentations --fast --preview
Note: TotalSegmentator only works with a NVidia GPU. If you do not have one you can try our online tool: www.totalsegmentator.com
--fast: For faster runtime and less memory requirements use this option. It will run a lower resolution model (3mm instead of 1.5mm).--preview: This will generate a 3D rendering of all classes, giving you a quick overview if the segmentation worked and where it failed (seepreview.pngin output directory).--statistics: This will generate a filestatistics.jsonwith volume (in mm³) and mean intensity of each class.--radiomics: This will generate a filestatistics_radiomics.jsonwith radiomics features of each class. You have to install pyradiomics to use this (pip install pyradiomics).
We also provide a docker container which can be used the following way
docker run --gpus 'device=0' --ipc=host -v /absolute/path/to/my/data/directory:/workspace wasserth/totalsegmentator_container:master TotalSegmentator -i /workspace/ct.nii.gz -o /workspace/segmentations
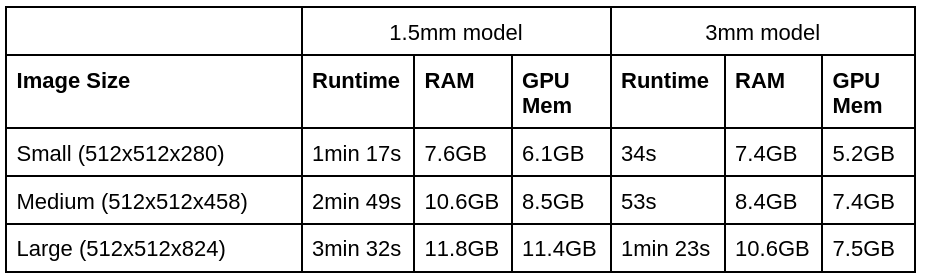
Totalsegmentator has the following runtime and memory requirements:
(1.5mm is the normal model and 3mm is the --fast model)
The exact split of the dataset can be found in the file meta.csv inside of the dataset. This was used for the validation in our paper.
The exact numbers of the results for the high resolution model (1.5mm) can be found here. The paper shown these numbers in the supplementary materials figure 11.
You have to download the data and then follow the instructions of nnU-Net how to train a nnU-Net. We trained a 3d_fullres model and the only adaptation to the default training is setting the number of epochs to 4000 and deactivating mirror data augmentation. The adapted trainer can be found here.
If you want to combine some subclasses (e.g. lung lobes) into one binary mask (e.g. entire lung) you can use the following command:
totalseg_combine_masks -i totalsegmentator_output_dir -o combined_mask.nii.gz -m lung
pip install git+https://github.com/wasserth/TotalSegmentator.git
For more details see this paper https://arxiv.org/abs/2208.05868. If you use this tool please cite it as follows
Wasserthal J., Meyer M., Breit H., Cyriac J., Yang S., Segeroth M. TotalSegmentator: robust segmentation of 104 anatomical structures in CT images, 2022. URL: https://arxiv.org/abs/2208.05868. arXiv: 2208.05868
Moreover, we would really appreciate if you let us know what you are using this tool for. You can also tell us what classes we should add in future releases. You can do so here.