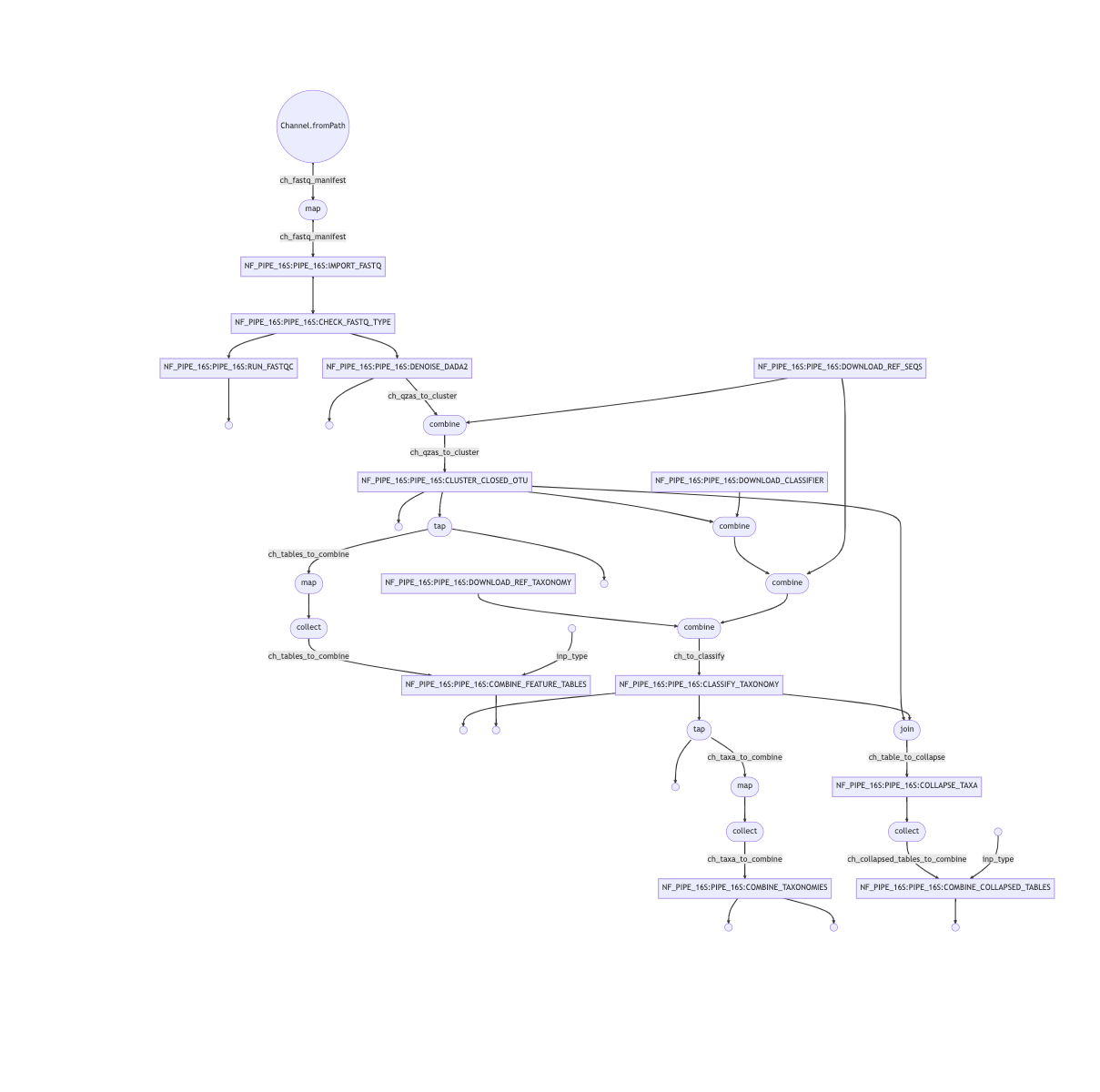
Workflow to process amplicon meta-analysis data, from either local FASTQs or NCBI accession IDs to taxonomic classification.
Note for users with newer Apple processors (M1/M2): Conda environments require emulation using Rosetta, due to the lack of certain packages for the ARM64 architecture otherwise available with Intel processors. Please follow the installation and setup instructions here for details.
Conda environments are available for all processes. To customize in run, modify the environment parameters (params.qiime_conda_env, params.fastqc_conda_env, params.multiqc_conda_env, and params.fondue_conda_env) in the input configuration file.
Launch a Conda environment-based run using -profile conda when running the workflow script.
Containers are available for all processes. To customize in run, modify the container parameters (params.qiime_container, params.fastqc_container, params.multiqc_container, and params.fondue_container) in the input configuration file.
Launch a container-based run with Singularity or Docker using -profile docker or -profile singularity when running the workflow script.
Unless otherwise noted, these parameters should be under the scope params in the run.config file.
Used for initial FASTQ processing.
read_type: FASTQ type, either"paired"or"single"
Required if running q2_fondue:
inp_id_file: Path to TSV file containing NCBI accession IDs for FASTQs to download. File must adhere to QIIME 2 metadata formatting requirements- Note: FASTQ file names starting with non-alphanumeric characters (particularly
#) are NOT supported. These will throw an error in your workflow!
- Note: FASTQ file names starting with non-alphanumeric characters (particularly
email_address: email address of user, required for SRA requests viaq2-fondue
Required if running from local FASTQ files:
fastq_manifest: Path to TSV file mapping sample identifiers to FASTQ absolute file paths; manifest must adhere to QIIME 2 FASTQ manifest formatting requirements
Required if running Cutadapt:
- For primer removal:
- If single-end,
cutadapt.front: Primer sequence to remove; multiple primers are delimited by a space. - If paired-end,
cutadapt.front_f/cutadapt.front_r: Primer sequences to remove; multiple primers are delimited by a space. - Note that
cutadapt.frontis recommended for most amplicon sequence runs, andcutadapt.adapterandcutadapt.anywhereare not supported in this workflow. The same goes for their paired counterparts. - The workflow does not at the moment support linked primers. Additionally, the workflow currently only takes a collection of single-end or paired-end primers, but not a combination of both.
- If single-end,
- To bypass the automated parameter validation, the user should set
params.validate_parameterstofalsewhen issuing the execution command.
-
By default, workflow inputs may be entered as TSV or FASTQ files; the workflow is designed to generate input QIIME 2 artifacts using the import/download processes. This behavior is controlled by the
generate_inputparameter, set totrueby default. -
To use an already-created input QIIME 2 artifact, the user should set
params.generate_inputtofalseand specify the path to the input artifact using theparams.input_artifactparameter. For example:
--generate_input false --input_artifact "path/to/input_artifact"Used for initial FASTQ processing in scope params.fastq_split:
enabled: defaultnull, determines whether samples will be processed as a batch or individually; either"True"or"False"method: default"sample", represents method by which to split input FASTQ file manifest; either"sample"or an integer representing the number of split artifacts for processingsuffix: default"_split.tsv", suffix for split FASTQ manifest files used as intermediates
Cutadapt process parameters in scope params.cutadapt:
num_cores: default1error_rate: default0.1indels: defaultTruetimes: default1overlap: default3, used for paired-end readsmatch_read_wildcards: default"False"match_adapter_wildcards: default"True"minimum_length: default1,discard_untrimmed: default"True"; we highly recommend keeping this parameter"True"as the Cutadapt process also separates reads by primer sequence!max_error_flag: defaultnullmax_n_flag: defaultnullquality_cutoff_5end: default0quality_cutoff_3end: default0quality_base: default33
DADA2 process parameters in scope params.dada2:
trunc_q: default2pooling_method: default"independent"chimera_method: default"consensus"min_fold_parent_over_abundance: default1.0num_threads: default0, to use all available cores on systemnum_reads_learn: default1000000hashed_feature_ids: default"True"- Parameters for single-end runs:
trunc_len: default0trim_left: default0max_ee: default2.0
- Parameters for paired-end runs:
trunc_len_f: default0trunc_len_r: default0trim_left_f: default0trim_left_r: default0max_ee_f: default2.0max_ee_r: default2.0min_overlap: default12
VSEARCH process parameters in scope params.vsearch:
perc_identity: default0.8strand: default"plus"num_threads: default0, to use a single thread per core
Feature classifier process parameters in scope params.classifier:
method: default"sklearn"; also accommodates"blast"and"vsearch"- Parameters for
sklearn-based classifier:reads_per_batch: default"auto"num_jobs: default-1pre_dispatch: default"2*n_jobs"confidence: default0.7read_orientation: default"auto"
- Parameters shared between BLAST+ and VSEARCH consensus classifiers:
max_accepts: default10perc_identity: default0.8query_cov: default0.8strand: default"both"min_consensusdefault0.51unassignable_label: default"Unassigned"
- Additional parameters for BLAST+ classifier:
evalue: default0.001
- Additional parameters for VSEARCH classifier:
search_exact: default"False"top_hits_only: default"False"max_hits: default"all"max_rejects: default"all"output_no_hits: default"True"weak_id: default0.0num_threads: default1
VSEARCH reference-based chimera identification process parameters in scope params.uchime_ref:
dn: default1.4min_diffs: default3min_div: default0.8min_h: default0.28xn: default8.0num_threads: default1
Additional process parameters:
taxa_level: default5, collapsing taxonomic classifications to genus; used inqiime taxa collapsephred_offset: default33; used in FASTQ import if using local FASTQsvsearch_chimera: default"False"
Reference files if available locally; otherwise, defaults will be downloaded from the QIIME 2 data resources page:
otu_ref_file: defaultnull, downloading pre-formatted files from the SILVA 138 SSURef NR99 full-length sequences; used in closed-reference OTU clustering with VSEARCHtrained_classifier: defaultnull, downloading naive Bayes taxonomic classifiers trained on SILVA 138 99% OTUs full-length sequences; used in taxonomy classificationtaxonomy_ref_file: defaultnull, downloading pre-formatted file from the SILVA 138 SSURef NR99 full-length taxonomy; used inq2-feature-classifierif running with BLAST+
For containerization:
qiime_release: default"2023.2", used to specify paramqiime_containerto particular QIIME versionqiime_container: default"quay.io/qiime2/core:${params.qiime_release}"; location of QIIME container used for workflow; if running on platforms without Internet, point to a valid .sif file. Note that local files must be prefixed withfile://; triple/denotes absolute filepaths.qiime_conda_env: default"${baseDir}/assets/qiime2-2023.2-py38-${sys_abbreviation}.yml"fastqc_release: default"v0.11.9_cv8", used to specify paramfastqc_containerto particular FastQC image versionfastqc_container: default"biocontainers:fastqc"; location of Docker container used for FastQC processesfastqc_conda_env: default"bioconda::fastqc"fondue_release: default"2023.2-ps", used to specify paramfondue_containerto particular q2-fondue image versionfondue_container: default"linathekim/q2-fondue:${fondue_release}"- Note: The standard environment for
q2-fonduewill not work with Nextflow out of the box. The image requires installation ofprocps(available withapt-get) for interactions with Nextflow. These are denoted with the suffix-pson DockerHub.
- Note: The standard environment for
fondue_conda_env: default"${baseDir}/assets/q2-fondue-2023.2-${sys-abbreviation}.yml"multiqc_release: default"v1.18"multiqc_container: default"ewels/multiqc:${multiqc_release}"multiqc_conda_env: default"bioconda::multiqc"
These run configurations fall under non-param scopes listed below.
Reporting with Nextflow Tower (scope tower):
enabled: defaultfalse, allowing workflow metrics to be reported in the Nextflow Tower interfaceaccessToken: user token for Nextflow Tower reporting; required if running Nextflow Tower, unlessTOWER_ACCESS_TOKENhas otherwise been defined in the runtime environment
Execution parameters (scope process):
executor: default"local", resource manager to run workflow on; options include"slurm","sge","awsbatch", and"google-lifesciences"withLabel:container_qiime2.container: default${params.qiime_container}, but can be replaced with location of local container containing QIIME 2 core distribution
To skip processes through DADA2, if using pre-denoised feature tables and sequences:
denoised_table: Path to QIIME 2 artifact containing a denoised feature tabledenoised_seqs: Path to QIIME 2 artifact containing denoised sequences corresponding with the above feature table
outDir/taxonomy.qza: Artifact containing frequencies for features collapsed to a given level (default genus).outDir/taxonomy.qzv: Visualization containing frequencies for features collapsed to a given level (default genus).outDir/feature_table.qza: Artifact containing table of represented features by sample.outDir/stats/: Directory containing QC metrics, including FastQC, clustering statistics, denoising statistics, etc.outDir/trace/: Directory containing runtime metrics with an execution report and a pipeline DAG.
- Data import (
qiime tools import) or FASTQ download (q2-fondue) - Optional adapter trimming:
q2-cutadapt - Initial quality control and denoising:
q2-dada2 - Optional chimera filtering:
q2-vsearch - Closed reference OTU clustering:
q2-vsearch - Taxonomy classification:
q2-feature-classifier - Collapse to taxon of interest and merge final outputs
nextflow run /path/to/workflow/main.nf -c run.config -profile conda