- Uniform data model, with ability to store arbitrary source-specific metadata.
- Multi-tenancy support using tenant-id.
- Scalable and extensible.
- Can add new sources.
- Can add new ingestor types.
- Can create/update ingestion pipeline.
- Can scale horizontally on multiple servers very easily.
- Multiple feedback sources of same source type for same tenant.
- Can enrich data ingested at any step of the pipeline.
This framework works on 2 interfaces: IngestionWriter and Ingestor.
IngestionWriterinterface defines aWritemethod which allows concrete implementations to write the data to a database, repository or forward data to anIngestor.Ingestorinterface defines aListenAndServemethod which allows concrete implementations to receive data, process it and handle it with help ofIngestionWriter.
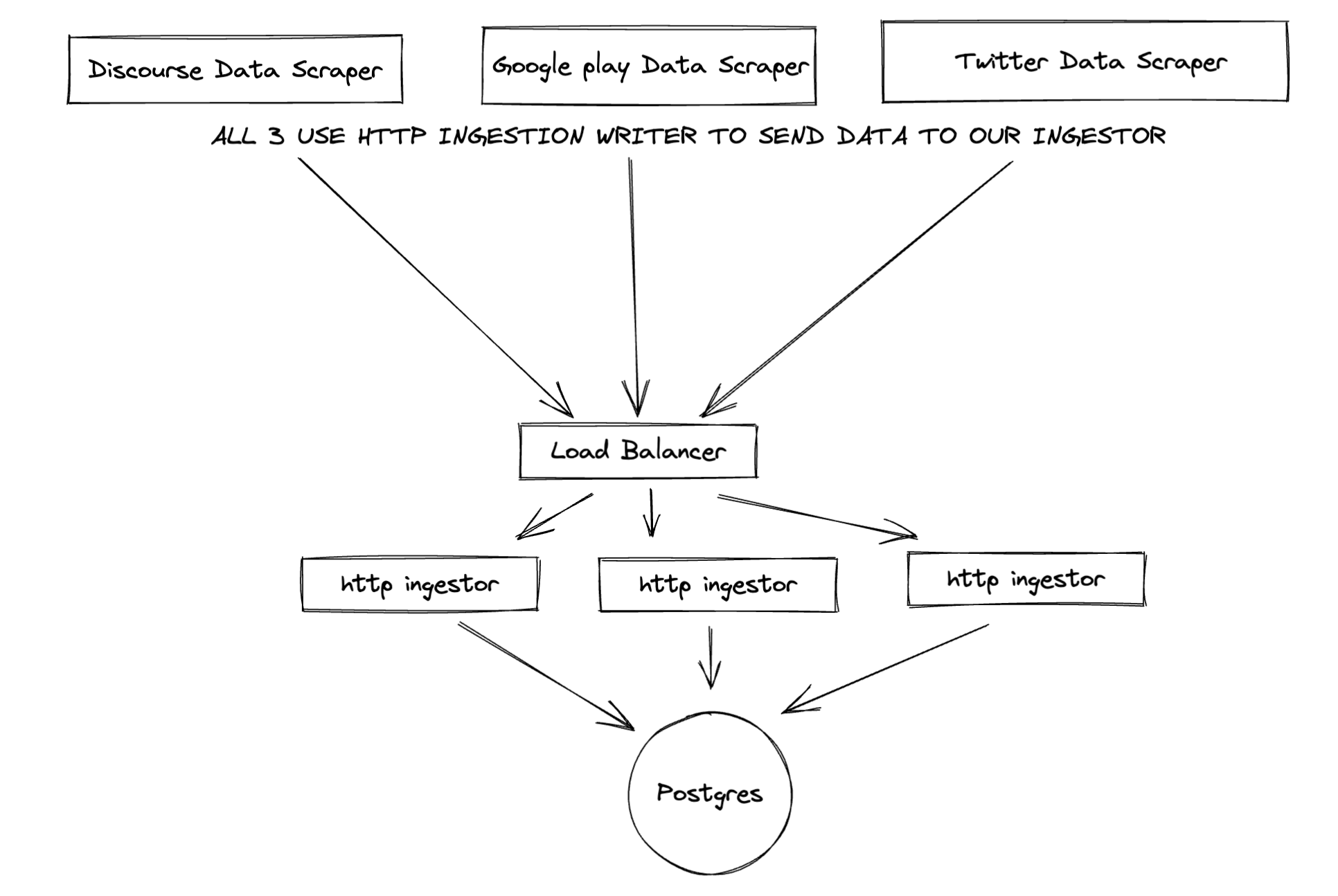
- Our data fetching services, example:
DiscourseScraperare composed with an implementation ofIngestionWriterinterface. That allows them the flexibility to switch Ingestion Writers if needed, without changing their own code, as well as code reuse. - If our DataFetchingServices use an
IngestionWriter(for example, seeSimpleHttpIngestionWriter) which sends data to anIngestor, which receives it, handles it appropriately and then sends it to its ownIngestionWriter. ThatIngestionWriter, depending on the concrete implementation can choose to save the data to a database / forward it to another ingestor based on custom logic. - Suppose we add another Ingestor, which uses GRPC, then we just need to create concrete implementions
GRPCIngestionWriterandGRPCIngestor, and we can switch them out with other implementations. - This way, we have abstracted out the communication, allowing us to focus on core logic of "Data fetching", "Data enrichment" etc.
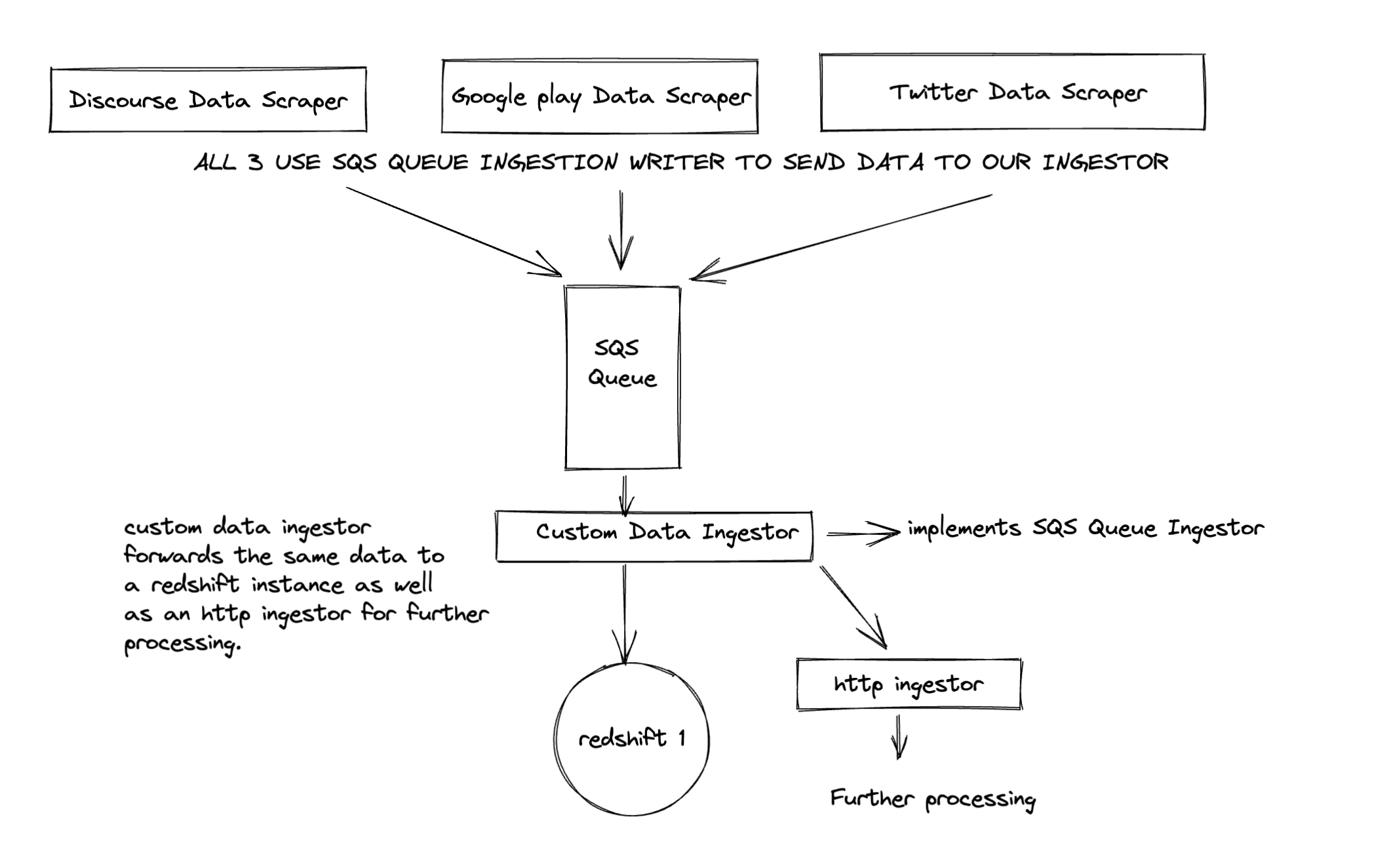
- A more complex use case is shown below, where different data fetching services publish to an SQS queue using
SQSIngestionWriter. TheSQSIngestorreceives it and a custom writer forwards the data to anHttpIngestoras well as writes the data toRedshift.
After the data is scraped/fetched, data can only be sent in our pipeline in format of IngestionData. However, all fields in IngestionData might not
be known/available to us at that time. In that scenario, known data can be filled and other data dumped in Data field as well as Metadata fields.
Our ingestion system can very easily adapt to this by having some preprocessing ingestors which ingest half filled data, enriches it by deriving/adding remaining attributes before passing it on to the main ingestion pipeline.
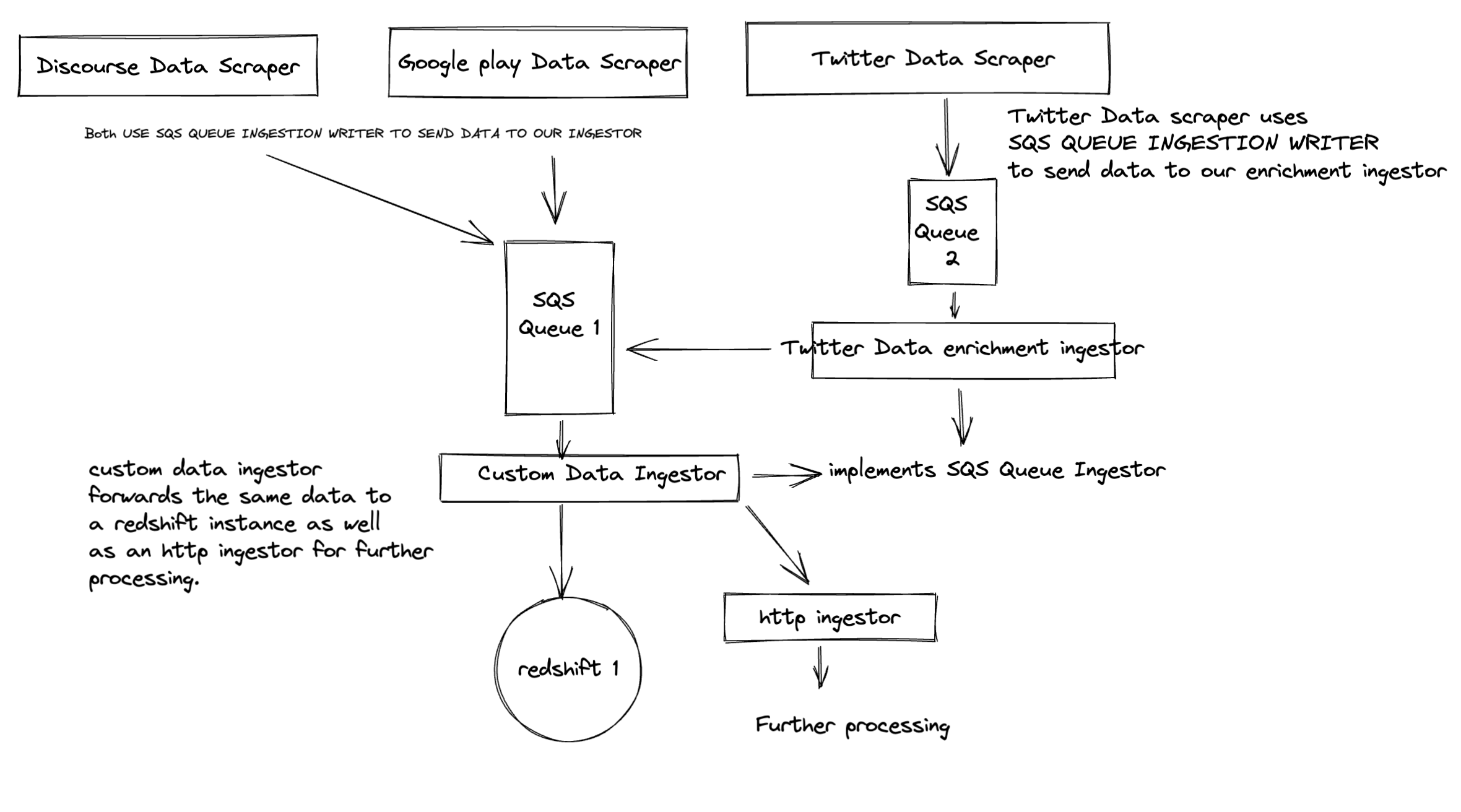
For example: Let's say a tenant has multiple apps, but only one twitter account for all the apps. When we fetch twitter data, it might not be possible to know which app the tweet is about. In that case, we can fix the above use case scenario like this:
It depends on the implementation of our Ingestor and IngestorWriter Interface, we support both. For example, in SQS, pull based mechanism is used.
For http, the data is being pushed to our ingestors. To me, using a msg queue like SQS makes the "perfect sense", so that ingestors can process data at their own speed. So pull based approach would work better in data intensive scenarios.
metadata attributes are always growing and all of them cannot be thought about at the initial stage. So it makes sense for it to be a custom JSON value with unknown attributes. Additionally, we add metadata_type as an additional attribute seperately, which lets us distinguish between different metadata types, to allow processing of data using metadata values present, or enriching of metadata using ingestors.
IngestionData has support for record level attributes. These record level attributes can be provided by our data fetching services, or an intermediate Ingestor in the pipeline can set them using different strategies.
Idempotency is an implmentation detail left to IngestionWriter. There are different types of cases in which idempotency might be needed. 2 of those are:
- An
IngestionWriterwriting a record to aIngestortimes out and retries. TheIngestionWriteronIngestorside needs to be implemented in a way that duplicate uuid ofIngestionDatadoesn't lead to duplicate records. This might not be needed when our ingestor is just doing some processing and forwarding the data to another ingestor, but is needed in cases when it is storing the data in a repository. Implementation is easy, a quickSELECTbeforeINSERTin SQL, for example. - An
IngestionWriteris forwarded with data already present in the database, but with a different UUID. This can happening cases when the server is restarted with a certain backfill time. This might not be needed in cases whenIngestoris just doing some processing and itsIngestionWriterforwarding the data to anotherIngestor, but in cases of saving data to a repository, a combined check of(source_id, source_type, data_id)in the repository will prevent us from adding duplicate records.
SourceIdfield inIngestionDataallows this
-
A demo in which random data is read and then forwarded to a
HttpIngestorhas been written. TheHttpIngestorwrites data to an postgres db usingPostgresIngestionWriter. To run it:cd examples/postgres-ingestion/scripts docker-compose build docker-compose upIn another terminal, from root directory
go run examples/mock-src/main.go
-
A demo in which limited data with few fields missing is available. An
HttpIngestorprocesses the data, adds new fields and sends it to ourHttpIngestorcreated in our earlier demo. To run it:# Make sure the final step in the ingestors is running, i.e # http ingestor with postgres writer cd examples/postgres-ingestion/scripts docker-compose build docker-compose up # Now, in new tab, from root dir, let's go to our data enrichment ingestion example. # Run the server using cd examples/data-enrichment-ingestion/scripts docker-compose build docker-compose up # In another terminal, from root directory, run the data generator go run examples/data-enrichment-ingestion/mock-src/main.go
PS: tests have not been written as the examples shown are non production ready POCs of actual implementation which should be done.