Antibody Analysis
Warning
This branch refers to our article:
Use the branch igpipeline2_timepoint_v2 if you are interested in the analyzes of the following articles:
Table of Contents
Installing Docker Desktop on macOS and Windows
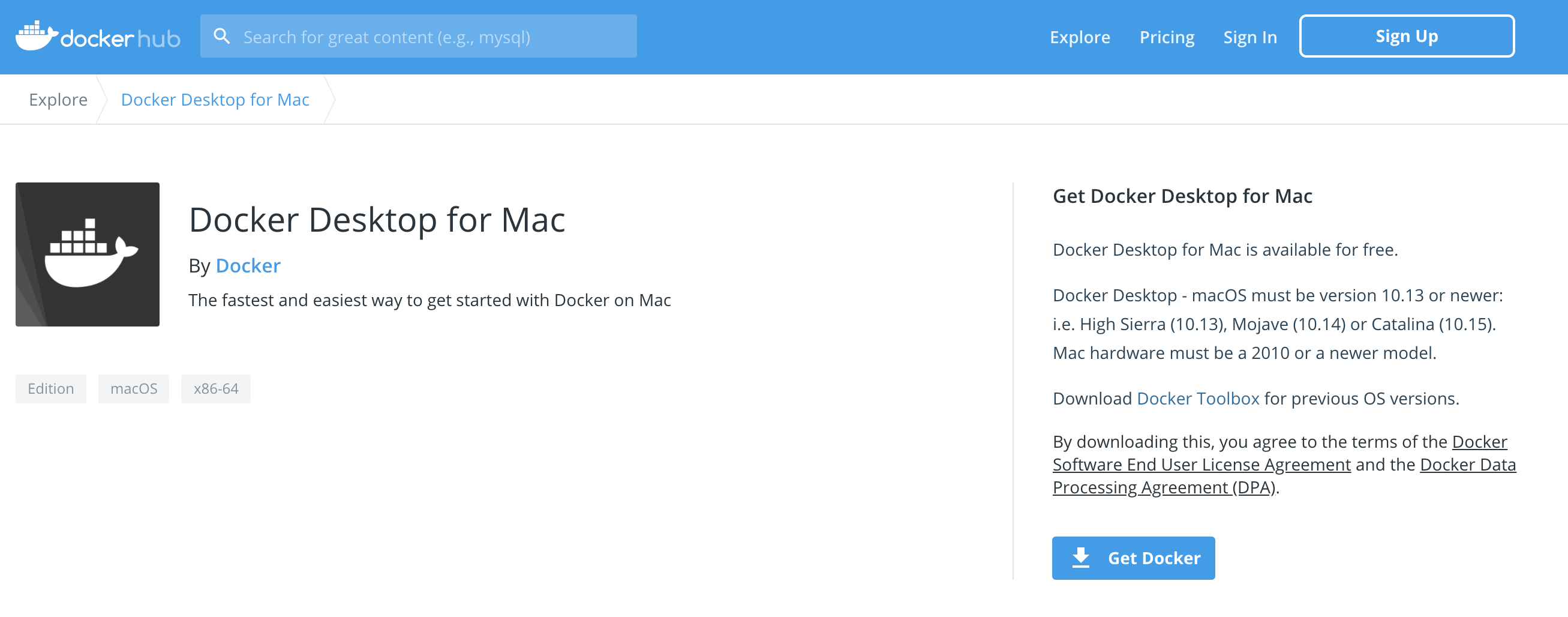
On the official Docker website, click on the button "Download for Mac" for macOS users or "Download for Windows" for Windows OS users.
Regardless the selected OS, a new web page will be open and the dmg file can be downloaded clicking on the button "Get Docker"
The same web page describes:
Install it on macOS
Double-click Docker.dmg to start the install process.
When the installation completes and Docker starts, the whale in the top status bar shows that Docker is running, and accessible from a terminal.

Install it on Windows
Double-click Docker for Windows Installer to run the installer.
When the installation finishes, Docker starts automatically. The whale  in the notification area indicates that Docker is running, and accessible from a terminal.
in the notification area indicates that Docker is running, and accessible from a terminal.
Executing IgPipeline
Step 1:
Once Docker is installed, download the image containing the IgPipeline on https://hub.docker.com/r/stratust/igpipeline
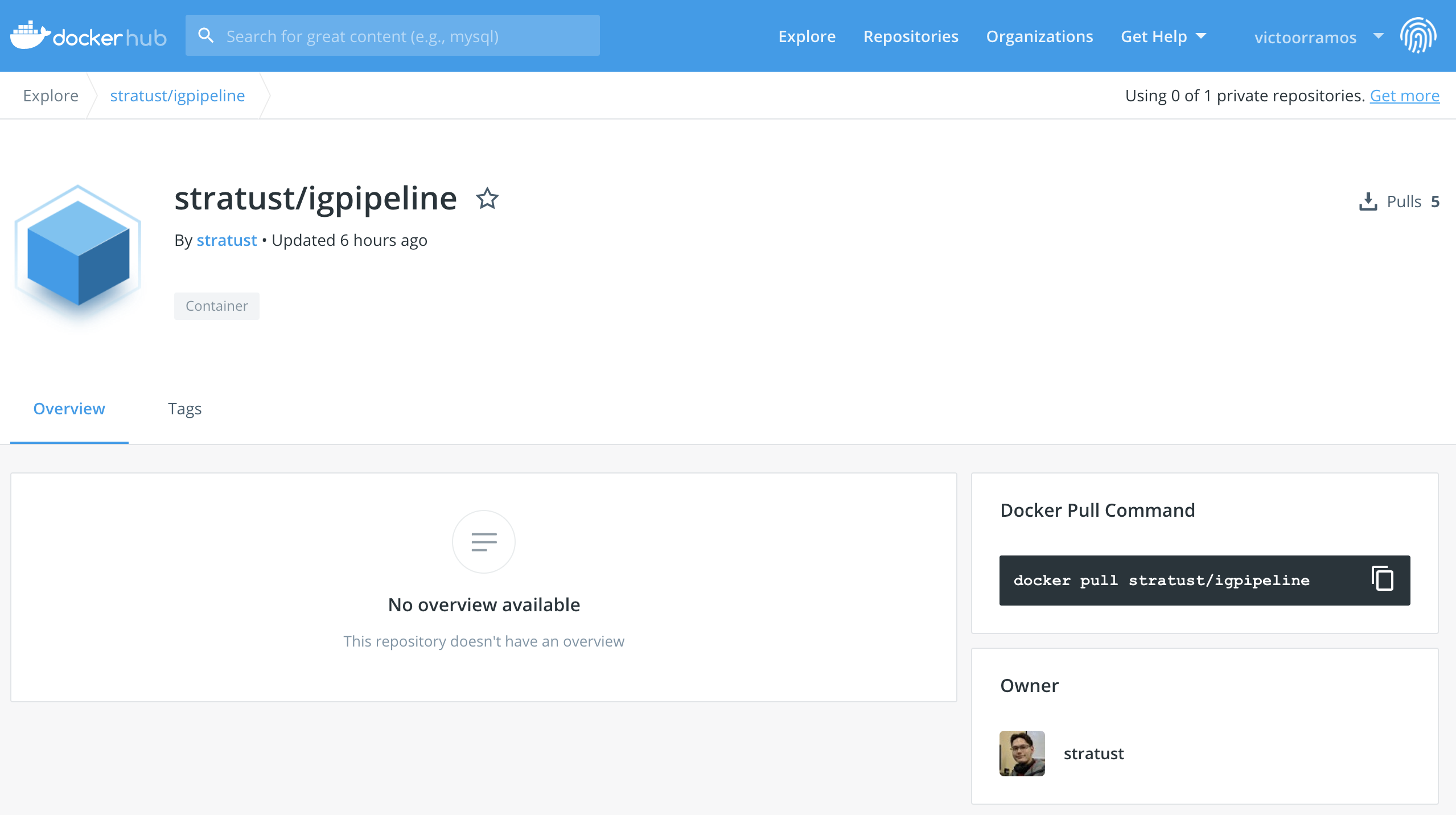
Open up a terminal session and download the image using the command docker pull stratust/igpipeline
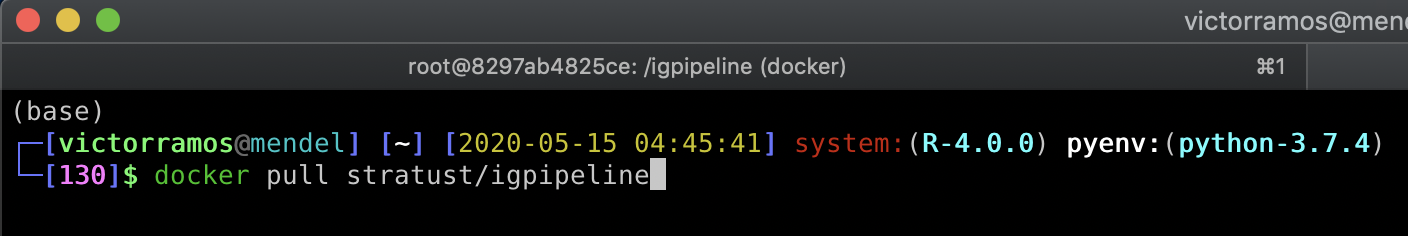
Step 2:
In Desktop, create a folder named "ig_analysis". Inside this folder, download and extract the zip file available through this link and create a folder named "results"

Step 3:
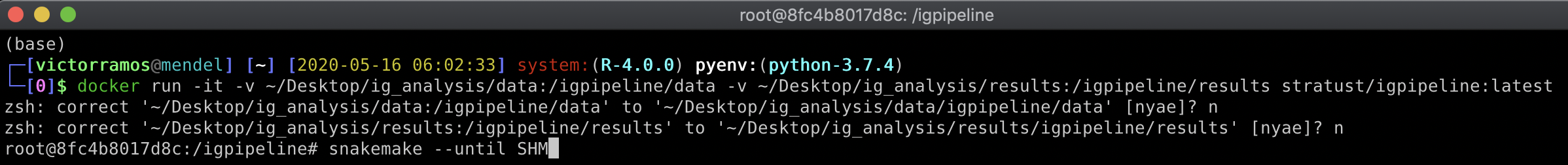
- To load a container with the downloaded image, open up a terminal session and type:
docker run -it -v ~/Desktop/ig_analysis/data:/igpipeline/data -v ~/Desktop/ig_analysis/results:/igpipeline/results stratust/igpipeline:latest
- Right after the container is loaded, to start the pipeline execution type:
snakemake --until SHM
Warnings
-
In the hydrophobicity analysis we calculate the GRAVY score for 22,654,256 IGH CDR3 sequences from a public database of memory B-cell receptor sequences (doi:10.1371/journal.pone.0160853), which requires plenty of computing resources and it takes ~30 min to execute. Using the command provided above this step WILL NOT be executed. If you want to execute the hydrophobicity score calculation, run the pipeline with the command snakemake
-
If you have enough computing resources to parallelize the execution, specify the parameter -j < number_of_cores > for the snakemake.
That's it ! The IgPipeline is executing.
Step 4:
Once the execution finishes, the results will be available in the folder "results"