An R package for implementing neural-g, a neural network-based approach for g-modeling, or mixing density estimation in a latent variable model.
Neural-g is a flexible neural network-based approach for estimating the mixing density in a latent variable model. Neural-g is capable of estimating a variety of latent densities, including atomic, smooth, flat, piecewise constant, and heavy-tailed densities. Neural-g works well under default hyperparameters that do not require additional tuning.
To run the neural-g smoothly, there are several prerequisites that need to be installed before using the R package. The main functions neural_g and neural_g2 are written in Python, namely the Pytorch library. We strongly recommend using CUDA (GPU-Based tool) to train neural-g, which offers magnitudes of speedup over using CPU.
- Python 3.7 or above
- Pytroch 1.11.0 or above
- NAVID CUDA 10.2 or above
In R, we also need reticulate package to run Python in R and devtools to install R package from github.
install.package("reticulate")
install.package("devtools")
Now, use the following code to install neuralG package.
library(devtools)
install_github(repo = "shijiew97/neuralG")
library(neuralG)
There are two main functions in the neuralG package, which are detailed below.
neural_ggives the neural-g estimator for a univariate mixture model. Currentlyneural_gsupports the following mixture models: Gaussian-location, Poisson-mixture, Lognormal-location, Gumbel-location, Gaussian-scale, Binomial-prob.neural_g2gives the bivariate neural-g estimator for a bivariate mixture model. Currentlyneural_g2supports Gaussian location-scale mixture model.
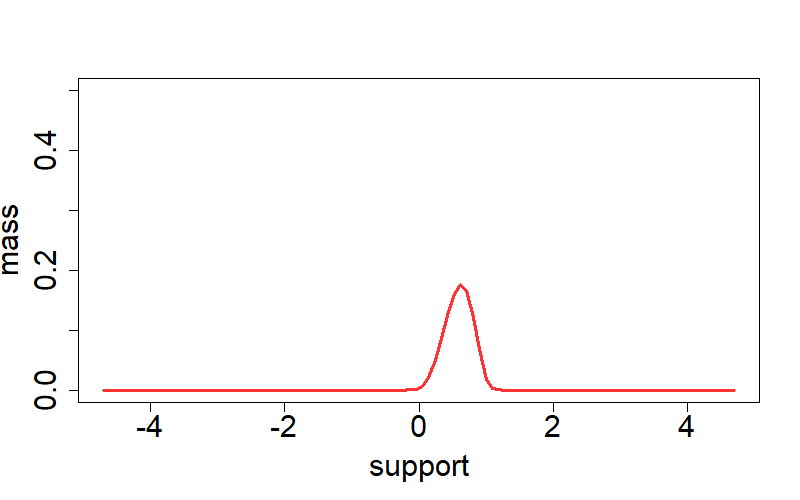
As a simple example of neuralG pacakge, we consider a Lognorm-location mixture:
Seed <- 128783;set.seed(Seed);dist <- "LogGaussian";param <- 0.2
n <- 2000;L <- 5;num_it <- 4000;n_grid <- 100
theta <- rbeta(n, 3, 2);Y = rlnorm(n, theta, param)
net_g <- neural_g(Y=Y, param=param, dist=dist, n=n, num_it=num_it, n_grid=n_grid)
plot(net_g$support, net_g$prob, col=rgb(1,0,0,0.8), type='l',
xlab='support', ylab='mass', lwd=3, ylim=c(0, 0.5), cex.axis=1.85, cex.lab=1.85)
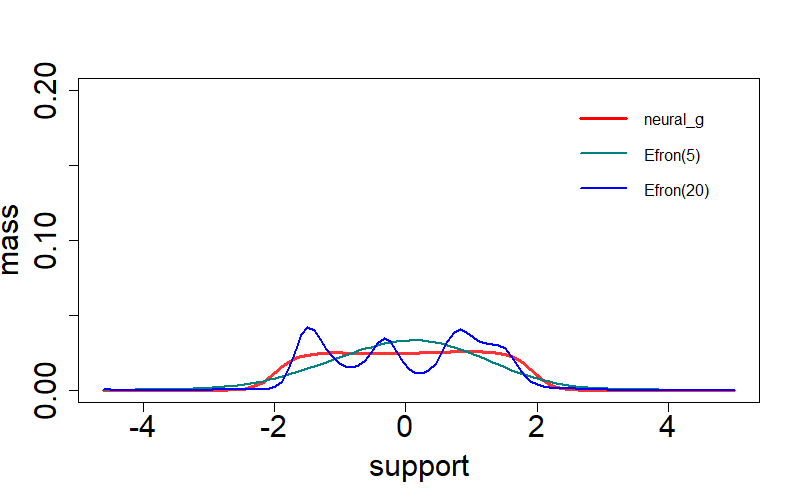
Here we also consider Efron's
Seed <- 128783;set.seed(Seed);dist <- "Gaussian";param <- 1.0
n <- 4000;L <- 5;num_it <- 4000;n_grid <- 100
theta <- runif(n, -2, 2);Y = theta + rnorm(n, 0, param)
net_g <- neural_g(Y=Y, param=param, dist=dist, n=n, num_it=num_it, n_grid=n_grid)
plot(net_g$support, net_g$prob, col=rgb(1,0,0,0.8), type='l',
xlab='support', ylab='mass', lwd=3, ylim=c(0,0.2), cex.axis=1.85, cex.lab=1.85)
efnpmle = deconvolveR::deconv(tau=net_g$support, X=Y, deltaAt=0, family="Normal", pDegree=5, c0=1.0)
efnpmle20 = deconvolveR::deconv(tau=net_g$support, X=Y, deltaAt=0, family="Normal", pDegree=20, c0=0.5)
lines(net_g$support, efnpmle$stats[,"g"], type="l", xlab="", ylab="", col=rgb(0,0.5,0.5), lty=1, lwd=2)
lines(net_g$support, efnpmle20$stats[,"g"], type="l", xlab="", ylab="", col=rgb(0,0,1), lty=1, lwd=2)
legend("topright", c("neural-g","Efron(5)","Efron(20)"),
col=c(rgb(1,0,0),rgb(0,0.5,0.5),rgb(0,0,1)), lty=c(1,1,1),
lwd=c(3,2,2,2,2), bty="n", cex=1.0, x.intersp=0.8, y.intersp=0.8)
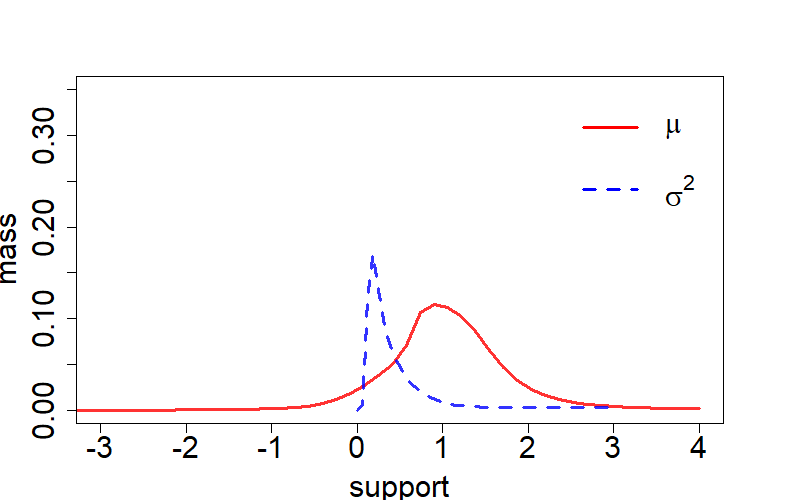
In this case, we explore the application of bivariate neural-g in a Gaussion location-scale mixture model:
Seed <- 128783;set.seed(Seed);dist <- "Gaussian2s";param <- 0.5
n <- 1000;L <- 5;num_it <- 4000;n_grid <- 50;p <- 2
mu <- 1;sigma <- 1; shape <- 2;scale <- 0.5
Y <- matrix(0, n, p)
theta1 <- MCMCpack::rinvgamma(n, shape, scale)
theta2 <- rnorm(n, mu, sd = sqrt(theta1 * sigma^2))
for(i in 1:n){Y[i,] <- theta2[i]+theta1[i]*rnorm(p)}
net_g <- neural_g2(Y=Y, param=param, dist=dist, n=n, num_it=num_it, n_grid=n_grid, p=p)
mu_support <- unique(net_g$support[,1])
sig_support <- unique(net_g$support[,2])
mu_prob <- rep(0, n_grid);sigma_prob <- rep(0, n_grid)
for(i in 1:n_grid){
mu_prob[i] <- sum( net_g$prob[net_g$support[,1]==mu_support[i]] )
sigma_prob[i] <- sum( net_g$prob[net_g$support[,2]==sig_support[i]] )
}
plot(mu_support, mu_prob, col=rgb(1,0,0,0.8), type='l', xlim=c(-3,4),
xlab='support', ylab='mass', lwd=3, ylim=c(0,0.35), cex.axis=1.85, cex.lab=1.85)
lines(sig_support, sigma_prob, col=rgb(0,0,1,0.8), type='l', lwd=3, lty=2)
legend(x=2.0, y=0.385, c(expression(mu),expression(sigma^2)),
col=c(rgb(1,0,0),rgb(0,0,1)), lty=c(1,2), lwd=c(3,3),
bty="n", x.intersp=0.5, y.intersp=0.8, seg.len=1.0, cex=1.85)