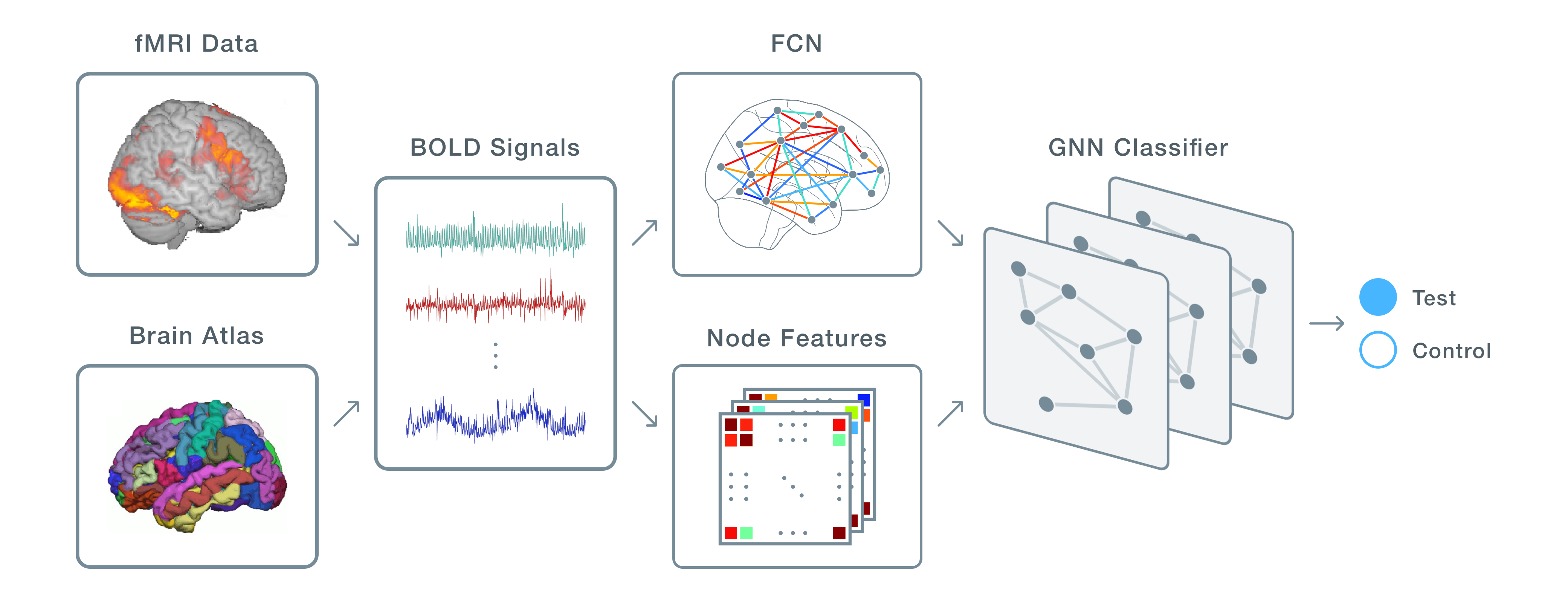
DeepFCN is a deep learning tool for predicting individual differences (e.g. classifying subjects with vs. without autism) from functional connectivity networks (FCNs).
It employs a Graph Neural Network (GNN) as a predictive model and offers control over every step in the machine learning pipeline, including:
- Extracting FCNs and features from fMRI data
- Preprocessing the FCNs (e.g. dropping outliers, normalization)
- Designing the GNN
- Training and testing the GNN
You can install DeepFCN from PyPi:
$ pip install deepfcnWe'll introduce the features of DeepFCN by applying it to the Autism Brain Imaging Data Exchange (ABIDE) dataset. Our goal is to train a model to predict whether a subject is diagnosed with autism.
We'll use the nilearn library to load the ABIDE dataset and a cortical brain atlas, which we'll use to parcellate the brain into regions of interest (ROIs):
import numpy as np
from nilearn.datasets import fetch_abide_pcp, fetch_coords_power_2011
from nilearn.input_data import NiftiSpheresMasker
autism_subjects = fetch_abide_pcp(DX_GROUP=1)["func_preproc"]
control_subjects = fetch_abide_pcp(DX_GROUP=2)["func_preproc"]
atlas = fetch_coords_power_2011()
coords = np.vstack((atlas.rois["x"], atlas.rois["y"], atlas.rois["z"])).T
roi_masker = NiftiSpheresMasker(seeds=coords, radius=5.0)The next step is to convert the subjects into data examples that can be submitted to a GNN. A single example is described by an instance of deepfcn.data.Example, which has following attributes:
node_features(numpy.ndarray): Node feature matrix with shape[num_nodes, num_node_features], wherenum_nodes == num_roisfc_matrix(numpy.ndarray): 3D functional connectivity matrix with shape[num_nodes, num_nodes, num_fc_features]; represents a multi-edge FCN, where each edge corresponds to a different measure of functional connectivity (e.g. correlation, covariance)y(int): Target to train against (e.g.0for autism,1for control)
To convert the ABIDE dataset into a set of examples, we'll use the deepfcn.data.create_examples function. This takes a set of fMRI scans (NiftiImage objects) as input and, for each scan, extracts the BOLD time series for each ROI, constructs an FCN, and extracts features to build an Example object. The method has the following parameters:
images(list): List of NiftiImage objects, each corresponding to a subject's fMRI scanlabel(int): Integer representing the target variable (i.e.y)roi_masker(NiftiMasker): Mask to apply when extracting time seriesfc_features(list, optional (default=["correlation"])): List of connectivity measures to use; options are listed in this tablenode_features(list, optional (default=["mean"])): List of node features to extract; options are listed in this tablen_jobs(int, optional (default=multiprocessing.cpu_count())): Number of CPUs to split up the work across
In our example, we'll use "correlation" and "dtw" (Dynamic Time Warping) as our connectivity measures, and "mean", "variance", and "entropy" as our node features:
from deepfcn.data import create_examples
fc_features = ["correlation", "dtw"]
node_features = ["mean", "variance", "entropy"]
examples = create_examples(autism_subjects, label=0, roi_masker=roi_masker,
fc_features=fc_features, node_features=node_features)
examples += create_examples(control_subjects, label=1, roi_masker=roi_masker,
fc_features=fc_features, node_features=node_features)If you wanted to define your own create_examples function, DeepFCN provides some helpers:
deepfcn.data.extract_signals(niimg, roi_masker): Extracts the BOLD time series for each ROIdeepfcn.data.extract_fcn(signals, feature_names): Creates a FCN from time series datadeepfcn.data.extract_node_features(signals, feature_names): Extracts node/ROI features from time series data
Now that we have a set of examples, the next step is to preprocess those examples before submitting them to a model. DeepFCN provides functions for cleaning graph-structured data, which we'll walk through below.
Because there's no "right" way to compare FCNs, there's also no right way to detect outliers in our dataset. However, DeepFCN still provides a technique for doing so, deepfcn.data.drop_outliers:
from deepfcn.data import drop_outliers
drop_outliers(examples, cutoff=0.05)This function does the following:
- Creates a vector representation of each example by averaging the node features and concatenating that with the mean of the edge features
- Calculates the Mahalanobis distance between each vector and the other example vectors
- Uses a Chi-Squared distribution to remove examples with distances outside a cutoff threshold (e.g. p < 0.05)
Within a FCN, there may be weak edges that represent noise rather than connectivity, and dropping them can improve performance of the GNN. deepfcn.data.drop_edges allows you to identify such edges and drop the ones below a connectivity threshold. If there are multiple connectivity measures associated with an edge, then only the first one is compared against the threshold. In our example, we'll drop the weakest 10% of edges from each example based on their correlation:
from deepfcn.data import drop_edges
drop_edges(examples, cutoff=0.10)Note that since some functional connectivity measures, such as correlation, are such that negative values are just as meaningful as positive values, only the absolute value is used –– e.g. a connectivity of -0.5 is considered stronger than 0.3.
Now that we've prepared our dataset, the next step is to create a GNN. This only takes one line of code to do:
from deepfcn.gnn import create_gnn
gnn = create_gnn(examples)By default, this function will autoconfigure the GNN parameters based on the number of node and edge features in the example set. It returns a PyTorch nn.Module object that leverages the NNConv and global_mean_pool functions from PyTorch Geometric, a graph classification library. If you want more control over how the GNN is configured, DeepFCN lets you tune various hyperparameters, listed below:
num_node_features(int): Number of node features in each input examplenum_edge_features(int): Number of edge features in each input examplehidden_channels(list): List of hidden channel sizes for each layer in the GNN; length of list corresponds to number of layers in the GNNuse_pooling(bool): Boolean indicating whether or not to use top k pooling after each layer in the GNNdropout_prob(float): Dropout probability to be applied to each GNN layerglobal_pooling_mode(str): Type of global pooling to use; options are"mean", forglobal_mean_pooling, and"attention", forGlobalAttentionconv_activation_func(nn.Module): Activation function to apply to the output of each graph convolution; must be a PyTorch activation functionedge_nn_kwargs(dict): TODOoutput_nn_kwargs(dict): TODO
These are all parameters in the deepfcn.gnn.create_gnn function.
DeepFCN offers a predefined and configurable training loop, deepfcn.gnn.cross_validate, to save you the trouble of creating your own. It has the following parameters:
examples(list): List of example objects (i.e. your dataset)gnn(nn.Module): PyTorch module representing the GNN to train and test (e.g. the output ofcreate_gnn)k(int): Number of folds to create for k-fold cross-validation
lr(float, optional (default=1e-3)): Learning rateepochs(int, optional (default=100)): Number of epochs to train forverbose(bool): IfTrue, training logs will be written tostdout
from deepfcn.gnn import cross_validate
results = cross_validate(examples, gnn, k=5, epochs=200)This function returns a list of dictionaries, each holding the cross-validation results for an epoch. The dictionaries have the following keys: "train_accuracy", "test_accuracy", "test_precision", "test_recall", "loss". Each key holds a list of k values, where each value corresponds to a fold used in cross-validation.
Because gnn is just a PyTorch module, you can also create your own training loop. Just note that doing this will require calling the to_data_obj() instance method on your deepfcn.data.Example objects to convert them into objects consumable by PyTorch Geometric modules.
TODO
| Feature | Description |
|---|---|
| correlation | Pearson correlation coefficient between the signals associated with two nodes/ROIs. |
| dtw | Speed-adjusted similarity between the two signals, calculated using the dynamic time warping algorithm. |
| granger_causality | Probability that activity in one node predicts the other. |
| efficiency | Multiplicative inverse of the shortest path distance between two nodes. Distance between two nodes is measured as the inverse of the absolute value of correlation. |
| Feature | Description |
|---|---|
| entropy | Complexity of the node's signal, based on approximate entropy of the time series. |
| fractal_dim | Complexity of the node's signal, based on the fractal dimension of the time series. |
| lyap_r | Largest Lyapunov exponent, calculated by applying the Rosenstein et al. algorithm to the time series. Positive exponents indicate chaos and unpredictability. |
| dfa | Measure of the "long-term memory" of a node's signal, computed using detrended fluctuation analysis. |
| mean | Mean of the node's signal. |
| median | Median of the node's signal. |
| range | Range of the node's signal. |
| std | Standard deviation of the node's signal. |
| auto_corr | Auto-correlation of the node's signal. |
| auto_cov | Auto-covariance of the node's signal. |
| weighted_degree | Weighted degree of the node, calculated by averaging the connectivity (correlation) of all its edges. |
| clustering_coef | Clustering coefficient for the node, where correlation is used as edge weight. |
| closeness_centrality | Reciprocal of the average shortest path distance to the node over all n-1 reachable nodes. Distance between two nodes is measured as the reciprocal of their connectivity (correlation). |
| betweenness_centrality | Sum of the fraction of all-pais shortest paths that pass through the node. |