Clustering Financial Time Series
This reporsitory contains the scripts and datasets used in my Master's Thesis on "Clustering Financial Time Series". The numerical experiments have been conducted using Python and R languages. One can find the necessary commands written below to reproduce the results in the thesis.
Python part
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import python_scripts.utils as tu
from python_scripts.hier_clust import *Here I illustrate the analysis on the Synthetic Dataset 1 and the same steps have been performed on Synthetic Dataset 2.
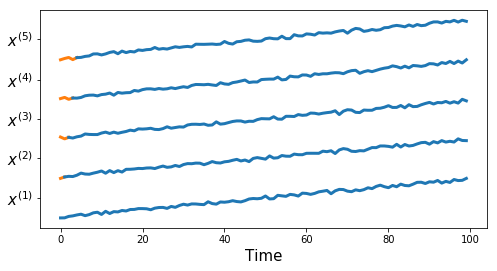
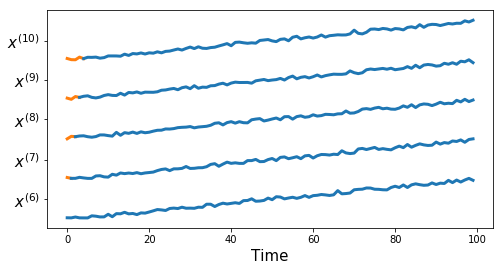
Synthetic Dataset 1
df, true_cluster = tu.get_synthetic_data('datasets/synthetic_data1.csv')
res_df, true_cluster = tu.get_synthetic_data('results/synthetic_data1_res.csv')plt.rcParams['figure.figsize'] = (8.0, 4.0)
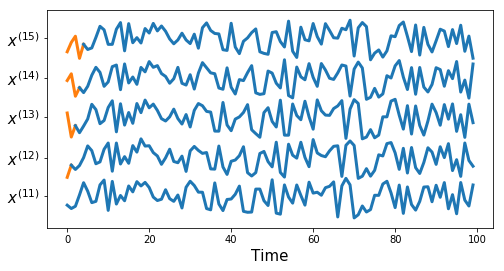
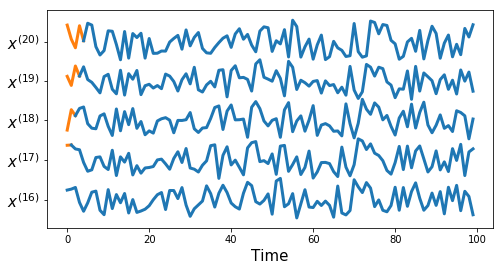
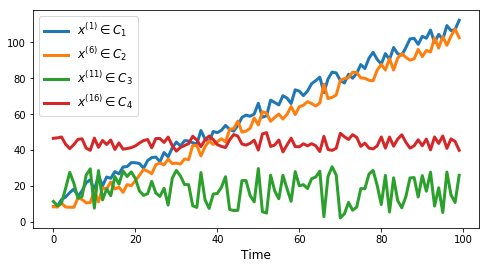
tu.plot_series(df, nr_series_per_class=5)tu.plot_one_per_class(df, nr_series_per_class=5)measures = ['euclidean', 'dtw', 'corr1', 'corr2',
'cross_corr1', 'cross_corr2', 'cross_corr3',
'acf', 'pacf', 'rccf1', 'rccf2', 'rccf3']get_sim_index(measures, df, res_df, true_cluster)| single | complete | average | |
|---|---|---|---|
| euclidean | 0.363636 | 0.380952 | 0.380952 |
| dtw | 0.380952 | 0.439935 | 0.380952 |
| corr1 | 0.334127 | 0.243590 | 0.294444 |
| corr2 | 0.334127 | 0.243590 | 0.243590 |
| cross_corr1 | 1.000000 | 1.000000 | 1.000000 |
| cross_corr2 | 1.000000 | 1.000000 | 1.000000 |
| cross_corr3 | 1.000000 | 1.000000 | 1.000000 |
| acf | 0.321429 | 0.321429 | 0.321429 |
| pacf | 0.363636 | 0.363636 | 0.363636 |
| rccf1 | 1.000000 | 1.000000 | 1.000000 |
| rccf2 | 1.000000 | 1.000000 | 1.000000 |
| rccf3 | 0.722222 | 0.949495 | 0.949495 |
chosen_measures = ['cross_corr2', 'cross_corr3',
'rccf2', 'rccf3']
cluster_numbers = list(range(2,11))get_sil_index(chosen_measures, cluster_numbers, df, res_df)| cross_corr2 | cross_corr3 | rccf2 | rccf3 | |
|---|---|---|---|---|
| 2 | 0.530485 | 0.739282 | 0.530762 | 0.692445 |
| 3 | 0.329060 | 0.426181 | 0.401696 | 0.477057 |
| 4 | 0.323237 | 0.420283 | 0.288671 | 0.288671 |
| 5 | 0.316615 | 0.413631 | 0.270898 | 0.270898 |
| 6 | 0.298892 | 0.404116 | 0.228582 | 0.228582 |
| 7 | 0.279041 | 0.384262 | 0.229174 | 0.229174 |
| 8 | 0.279625 | 0.384827 | 0.225471 | 0.225471 |
| 9 | 0.282080 | 0.387268 | 0.184505 | 0.184505 |
| 10 | 0.095913 | 0.095913 | 0.186782 | 0.186782 |
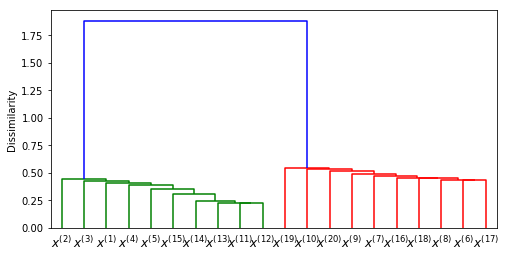
clustering = HClust(df, true_cluster, 'cross_corr3')
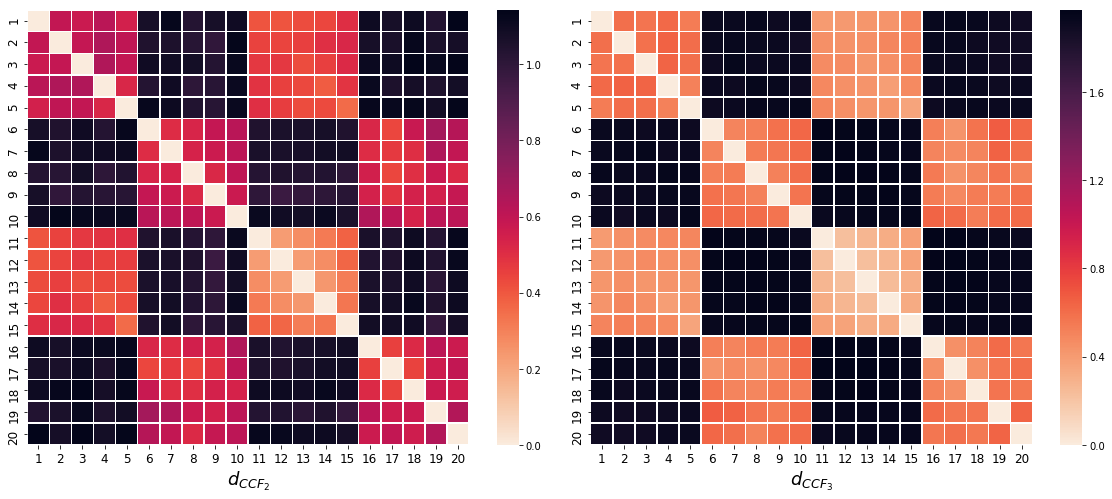
clustering.plot_dendrogram()clustering1 = HClust(df, true_cluster, 'cross_corr2')
clustering2 = HClust(df, true_cluster, 'cross_corr3')
fig = plt.figure(figsize=(20, 8))
fig.subplots_adjust(hspace=0.4, wspace=0.02)
fig.add_subplot(1, 2, 1)
clustering1.plot_heatmap(xlab='$d_{CCF_2}$')
fig.add_subplot(1, 2, 2)
clustering2.plot_heatmap(xlab='$d_{CCF_3}$')Clustering Stock Prices
stock_data = pd.read_csv('datasets/nyse_data.csv, index_col=0)
clustering = HClust(data=stock_data, ground_truth=None,
dist_func='cross_corr3', verbose=True)
clustering.dist_mat.to_csv('results/diss_mat_ccf3.csv', index=None, header=None)Dissimilarity computation: 100% [-------------------------------] Time: 0:32:53
diss_mat_ccf2 = pd.read_csv('results/diss_mat_ccf2.csv', header=None, index_col=None)
diss_mat_ccf3 = pd.read_csv('results/diss_mat_ccf3.csv', header=None, index_col=None)diss_ccf2 = np.triu(diss_mat_ccf2.values,1).flatten()
diss_ccf3 = np.triu(diss_mat_ccf3.values,1).flatten()
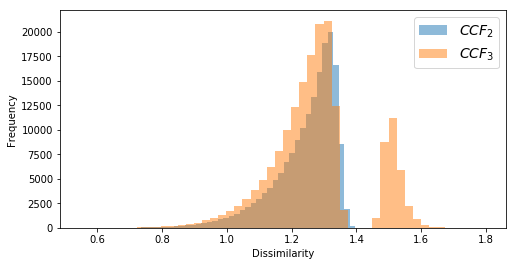
plt.hist(diss_ccf2[diss_ccf2!=0], bins=50, alpha=0.5, label='${CCF_2}$')
plt.hist(diss_ccf3[diss_ccf3!=0], bins=50, alpha=0.5, label='${CCF_3}$')
plt.xlabel('Dissimilarity')
plt.ylabel('Frequency')
plt.legend(fontsize=14);sectors = pd.read_csv('datasets/sectors.csv', header=None)Converting sector categories into numeric values.
from sklearn.preprocessing import LabelEncoder
sectors_numeric = LabelEncoder().fit_transform(sectors)get_similarities(diss_mat_ccf2, ground_truth=sectors_numeric)| single | complete | average | |
|---|---|---|---|
| 0.087304 | 0.325193 | 0.145864 |
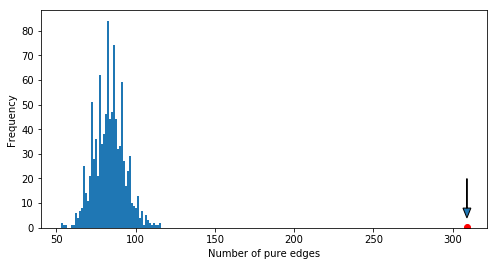
permutations = pd.read_csv('results/null_distribution1.csv')
permutations.plot(kind='hist', bins=50, legend=False)
s0 = 309
plt.plot(s0, 0.5, 'ro')
plt.arrow(s0, 20, 0, -16, length_includes_head=True,
head_width=5, head_length=4)
plt.xlabel('Number of pure edges');R part
source('R_scripts/clustering.R')
source('R_scripts/mst.R')df <- read.csv('datasets/synthetic_data1.csv')
nr.series <- 5
true_cluster <- c(rep(1, nr.series), rep(2,nr.series),
rep(1, nr.series), rep(2, nr.series))
This snippet will return the similarity index of the clustering using Piccollo and Maharaj distances
with single linkage method. Optionally one can plot the resulting dendrograms by setting plot=TRUE.
Here I have used the TSclust package.
cluster_eval(df, dist.method='AR.PIC',
linkage.method = 'single',
true_cluster, plot=F)
cluster_eval(df, dist.method='AR.MAH',
linkage.method = 'single',
true_cluster, plot=F)
This snippet fits an AR or ARIMA(p,1,0) type of model on each time series in the datasets and returns the model residuals for RCCF dissimilarity measure.
res.df <- get.residuals(df)
write.csv(res.df, file="results/synthetic_data1_res.csv",
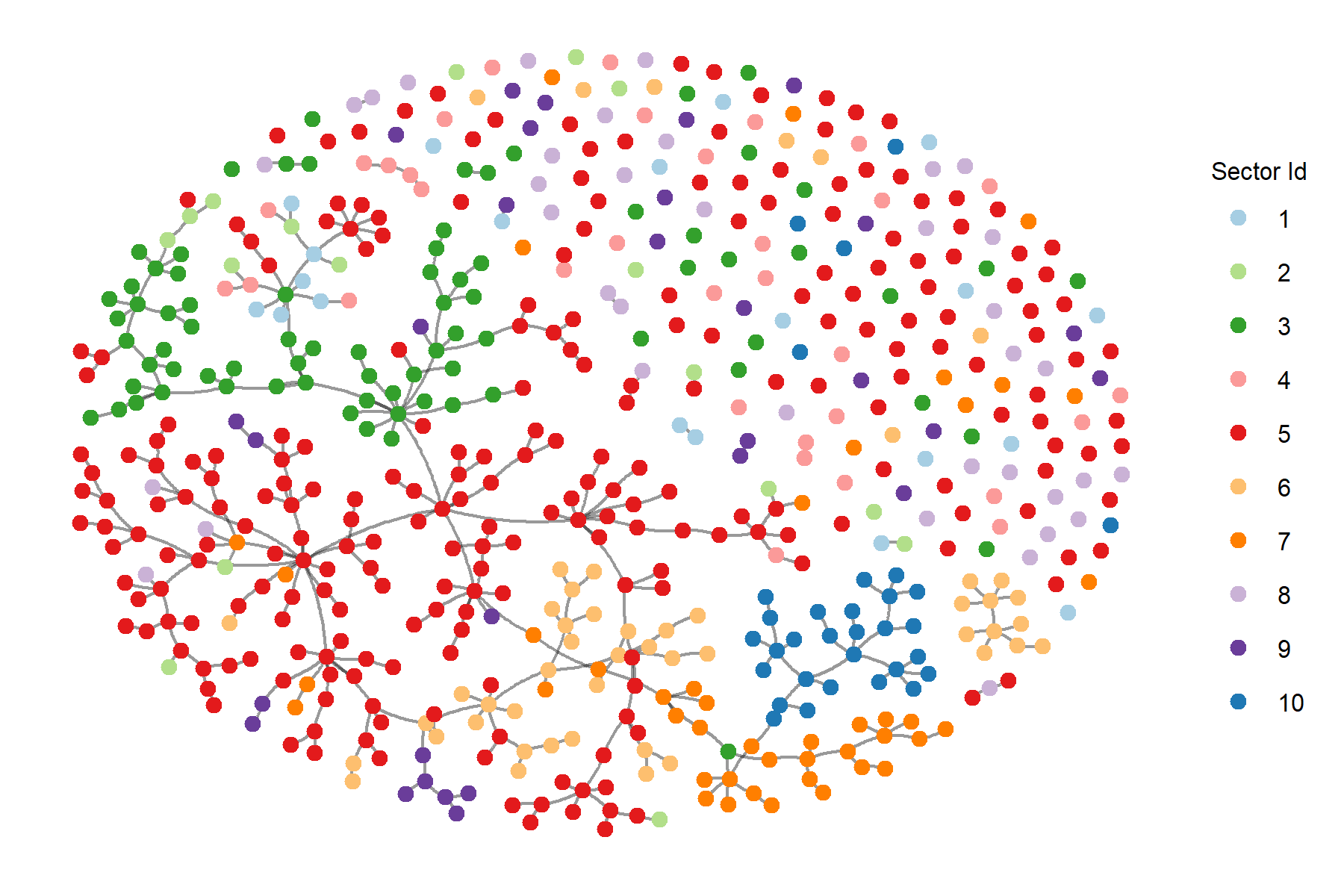
row.names = F)Plotting the minimum spanning tree with threshold 1 on the edge weights.
diss_data <- read.csv('results/diss_mat_ccf3.csv', header=F)
sectors <- read.csv('datasets/sectors.csv', header=F)
sectors <- sectors$V1
sectors.numeric <- as.numeric(sectors)
gr <- graph.adjacency(as.matrix(diss_data),
mode='undirected',
weighted = T)
mstree <- igraph::mst(gr)
final_mstree <- plot.mst(mstree, names=sectors.numeric,
pallete='Paired',
threshold=1,
save.fig = F,
fig.size=c(6,4))Running the permutation test will result in the p-value of the test and the resulting permutations will be saved as a csv file.
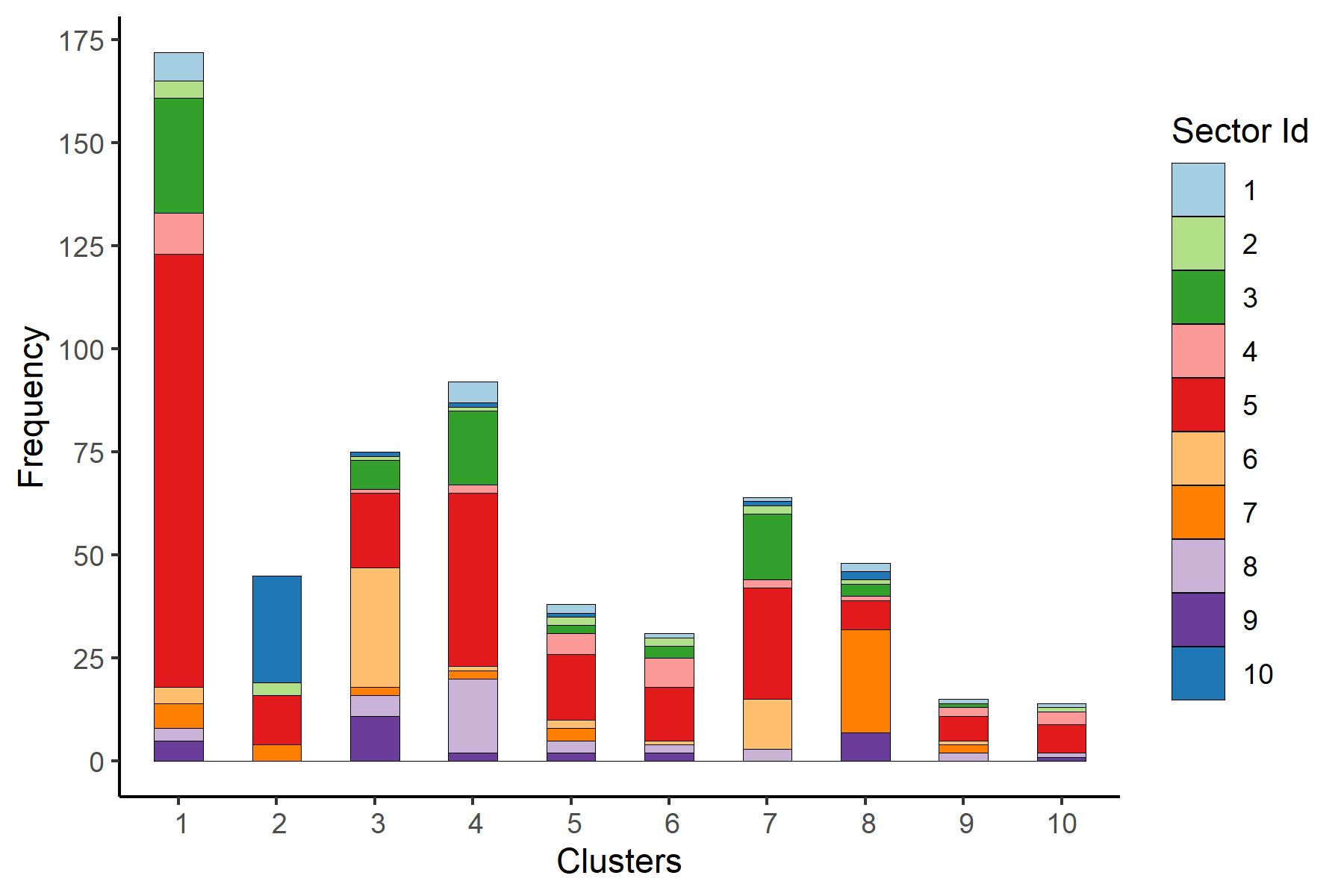
permutation.test(final_mstree, 10^4)Plotting the histogram of clusters in order to see the alignment between the obtained clusters and the provided categories (sectors).
df <- read.csv('results/clusters_complete_3.csv')
df$clusters <- df$clusters + 1
df$gt_num <- as.character(as.numeric(df$gt))
plot.hist(df, save.fig=T, fig.size=c(6,4))