This is a tutorial package that familiarizes intermediate R users with aspects of Bioconductor and introduces them to the Bioconductor package ecosystem.
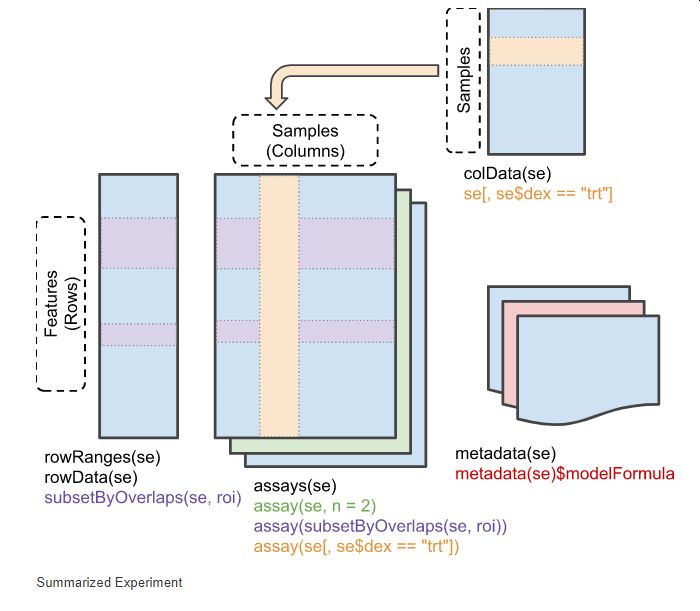
The SummarizedExperiment class is one of the fundamental ways to store data in Bioconductor, whether it be protein expression, RNASeq data, or single cell data. SummarizedExperiment contains information about a studies' experimental design, and information about the genes or entities that the assay data maps to.
- Learn about Bioconductor objects and how they enable complex analysis
- Learn about the components of a
SummarizedExperimentobject and how they work together - Extract experimental information about a
SummarizedExperimentobject usingmetadata() - Learn and Utilize sample information about the experimental design of a
SummarizedExperimentusingcolData() - Learn and utilize genomic information in
rowData()to select rows based on genomic coordinates - Construct complex subsetting queries using
colData()(selecting samples based on experimental design) androwData()(selecting reads within a Genomic range usingsubsetByOverlap()) - Utilize the
Deseq2package to run differential expression analysis using aSummarizedExperimentand a experimental design (specified by a formula)
install.packages("remotes")
remotes::install_github("laderast/biocowl")
library(biocowl)
learn_summarized_experiment()
Based on discussions with Kate Hertweck about how to teach Bioconductor. Parts of this tutorial are derived from https://bioconductor.org/packages/release/bioc/vignettes/SummarizedExperiment/inst/doc/SummarizedExperiment.html
Please note that the biocowl project is released with a Contributor Code of Conduct. By contributing to this project, you agree to abide by its terms.