Fit Gamma-Poisson Generalized Linear Models Reliably.
The core design aims of gmlGamPoi are:
- Fit the Gamma-Poisson models on arbitrarily large or small datasets
- Be faster than alternative methods, such as
DESeq2oredgeR - Calculate exact or approximate results based on user preference
- Support in memory or on-disk data
- Follow established conventions around tools for RNA-seq analysis
- Present a simple user-interface
- Avoid unnecessary dependencies
- Make integration into other tools easy
You can install the release version of glmGamPoi from BioConductor:
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("glmGamPoi")For the latest developments, see the GitHub repo.
Load the glmGamPoi package
library(glmGamPoi)To fit a single Gamma-Poisson GLM do:
# overdispersion = 1/size
counts <- rnbinom(n = 10, mu = 5, size = 1/0.7)
# design = ~ 1 means that an intercept-only model is fit
fit <- glm_gp(counts, design = ~ 1)
fit
#> glmGamPoiFit object:
#> The data had 1 rows and 10 columns.
#> A model with 1 coefficient was fitted.
# Internally fit is just a list:
as.list(fit)[1:2]
#> $Beta
#> Intercept
#> [1,] 1.504077
#>
#> $overdispersions
#> [1] 0.3792855The glm_gp() function returns a list with the results of the fit. Most
importantly, it contains the estimates for the coefficients β and the
overdispersion.
Fitting repeated Gamma-Poisson GLMs for each gene of a single cell dataset is just as easy:
I will first load an example dataset using the TENxPBMCData package.
The dataset has 33,000 genes and 4340 cells. It takes roughly 1.5
minutes to fit the Gamma-Poisson model on the full dataset. For
demonstration purposes, I will subset the dataset to 300 genes, but keep
the 4340 cells:
library(SummarizedExperiment)
library(DelayedMatrixStats)# The full dataset with 33,000 genes and 4340 cells
# The first time this is run, it will download the data
pbmcs <- TENxPBMCData::TENxPBMCData("pbmc4k")
#> snapshotDate(): 2020-04-27
#> see ?TENxPBMCData and browseVignettes('TENxPBMCData') for documentation
#> loading from cache
# I want genes where at least some counts are non-zero
non_empty_rows <- which(rowSums2(assay(pbmcs)) > 0)
pbmcs_subset <- pbmcs[sample(non_empty_rows, 300), ]
pbmcs_subset
#> class: SingleCellExperiment
#> dim: 300 4340
#> metadata(0):
#> assays(1): counts
#> rownames(300): ENSG00000126457 ENSG00000109832 ... ENSG00000143819
#> ENSG00000188243
#> rowData names(3): ENSEMBL_ID Symbol_TENx Symbol
#> colnames: NULL
#> colData names(11): Sample Barcode ... Individual Date_published
#> reducedDimNames(0):
#> altExpNames(0):I call glm_gp() to fit one GLM model for each gene and force the
calculation to happen in memory.
fit <- glm_gp(pbmcs_subset, on_disk = FALSE)
summary(fit)
#> glmGamPoiFit object:
#> The data had 300 rows and 4340 columns.
#> A model with 1 coefficient was fitted.
#> The design formula is: Y~1
#>
#> Beta:
#> Min 1st Qu. Median 3rd Qu. Max
#> Intercept -8.38 -6.43 -3.77 -2.46 1.01
#>
#> deviance:
#> Min 1st Qu. Median 3rd Qu. Max
#> 16.5 89.3 681 1695 5750
#>
#> overdispersion:
#> Min 1st Qu. Median 3rd Qu. Max
#> 0 0 0.388 1.15 15724
#>
#> Shrunken quasi-likelihood overdispersion:
#> Min 1st Qu. Median 3rd Qu. Max
#> 0.739 0.994 1 1.04 7.89
#>
#> size_factors:
#> Min 1st Qu. Median 3rd Qu. Max
#> 0.402 0.969 1 1.05 1.75
#>
#> Mu:
#> Min 1st Qu. Median 3rd Qu. Max
#> 9.24e-05 0.00158 0.0229 0.0871 4.8I compare my method (in-memory and on-disk) with DESeq2 and edgeR. Both are classical methods for analyzing RNA-Seq datasets and have been around for almost 10 years. Note that both tools can do a lot more than just fitting the Gamma-Poisson model, so this benchmark only serves to give a general impression of the performance.
# Explicitly realize count matrix in memory so that it is a fair comparison
pbmcs_subset <- as.matrix(assay(pbmcs_subset))
model_matrix <- matrix(1, nrow = ncol(pbmcs_subset))
bench::mark(
glmGamPoi_in_memory = {
glm_gp(pbmcs_subset, design = model_matrix, on_disk = FALSE)
}, glmGamPoi_on_disk = {
glm_gp(pbmcs_subset, design = model_matrix, on_disk = TRUE)
}, DESeq2 = suppressMessages({
dds <- DESeq2::DESeqDataSetFromMatrix(pbmcs_subset,
colData = data.frame(name = seq_len(4340)),
design = ~ 1)
dds <- DESeq2::estimateSizeFactors(dds, "poscounts")
dds <- DESeq2::estimateDispersions(dds, quiet = TRUE)
dds <- DESeq2::nbinomWaldTest(dds, minmu = 1e-6)
}), edgeR = {
edgeR_data <- edgeR::DGEList(pbmcs_subset)
edgeR_data <- edgeR::calcNormFactors(edgeR_data)
edgeR_data <- edgeR::estimateDisp(edgeR_data, model_matrix)
edgeR_fit <- edgeR::glmFit(edgeR_data, design = model_matrix)
}, check = FALSE, min_iterations = 3
)
#> # A tibble: 4 x 6
#> expression min median `itr/sec` mem_alloc `gc/sec`
#> <bch:expr> <bch:tm> <bch:tm> <dbl> <bch:byt> <dbl>
#> 1 glmGamPoi_in_memory 875.14ms 917.36ms 0.884 462.41MB 2.95
#> 2 glmGamPoi_on_disk 3.92s 4.06s 0.243 839.92MB 1.37
#> 3 DESeq2 19.95s 20.48s 0.0489 1.15GB 0.424
#> 4 edgeR 5.48s 6.12s 0.169 1.19GB 0.843On this dataset, glmGamPoi is more than 6 times faster than edgeR
and more than 20 times faster than DESeq2. glmGamPoi does not
use approximations to achieve this performance increase. The performance
comes from an optimized algorithm for inferring the overdispersion for
each gene. It is tuned for datasets typically encountered in single
RNA-seq with many samples and many small counts, by avoiding duplicate
calculations.
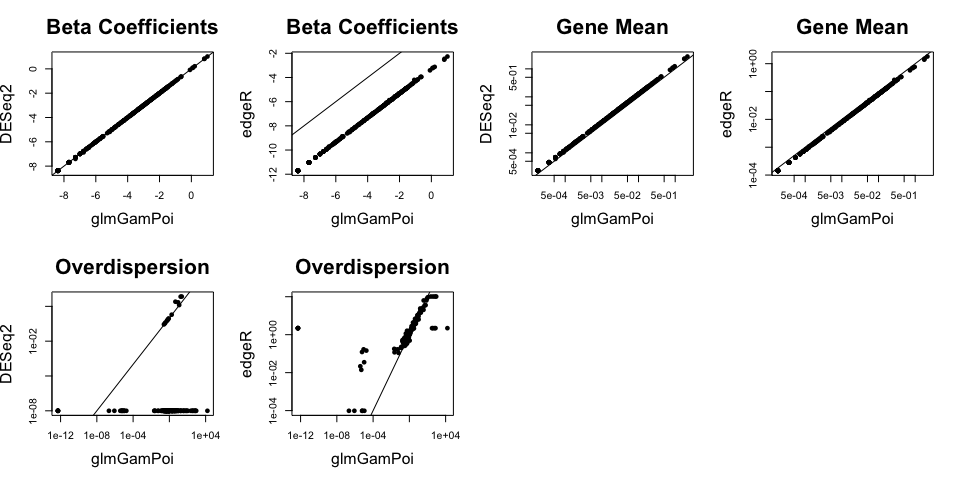
To demonstrate that the method does not sacrifice accuracy, I compare
the parameters that each method estimates. The means and β coefficients
are identical, but that the overdispersion estimates from glmGamPoi
are more reliable:
# Results with my method
fit <- glm_gp(pbmcs_subset, design = model_matrix, on_disk = FALSE)
# DESeq2
dds <- DESeq2::DESeqDataSetFromMatrix(pbmcs_subset,
colData = data.frame(name = seq_len(4340)),
design = ~ 1)
dds <- DESeq2::estimateSizeFactors(dds, "poscounts")
dds <- DESeq2::estimateDispersions(dds, quiet = TRUE)
dds <- DESeq2::nbinomWaldTest(dds, minmu = 1e-6)
#edgeR
edgeR_data <- edgeR::DGEList(pbmcs_subset)
edgeR_data <- edgeR::calcNormFactors(edgeR_data)
edgeR_data <- edgeR::estimateDisp(edgeR_data, model_matrix)
edgeR_fit <- edgeR::glmFit(edgeR_data, design = model_matrix)I am comparing the gene-wise estimates of the coefficients from all
three methods. Points on the diagonal line are identical. The inferred
Beta coefficients and gene means agree well between the methods, however
the overdispersion differs quite a bit. DESeq2 has problems estimating
most of the overdispersions and sets them to 1e-8. edgeR only
approximates the overdispersions which explains the variation around the
overdispersions calculated with glmGamPoi.
The method scales linearly, with the number of rows and columns in the
dataset. For example: fitting the full pbmc4k dataset with subsampling
on a modern MacBook Pro in-memory takes ~1 minute and on-disk a little
over 4 minutes. Fitting the pbmc68k (17x the size) takes ~73 minutes
(17x the time) on-disk. Fitting that dataset in-memory is not possible
because it is just too big: the maximum in-memory matrix size is 2^31-1 ≈ 2.1e9 is elements, the pbmc68k dataset however has nearly 100
million elements more than that.
glmGamPoi provides an interface to do quasi-likelihood ratio testing
to identify differentially expressed genes:
# Create random categorical assignment to demonstrate DE
group <- sample(c("Group1", "Group2"), size = ncol(pbmcs_subset), replace = TRUE)
# Fit model with group vector as design
fit <- glm_gp(pbmcs_subset, design = group)
# Compare against model without group
res <- test_de(fit, reduced_design = ~ 1)
# Look at first 6 genes
head(res)
#> name pval adj_pval f_statistic df1 df2 lfc
#> 1 ENSG00000126457 0.4540540 0.8928885 0.56061073 1 4431.149 NA
#> 2 ENSG00000109832 0.5836619 0.9151704 0.30039595 1 4431.149 NA
#> 3 ENSG00000237339 0.2572395 0.8123352 1.28388371 1 4431.149 NA
#> 4 ENSG00000075234 0.3812221 0.8776658 0.76691013 1 4431.149 NA
#> 5 ENSG00000161057 0.8507729 0.9924364 0.03539793 1 4431.149 NA
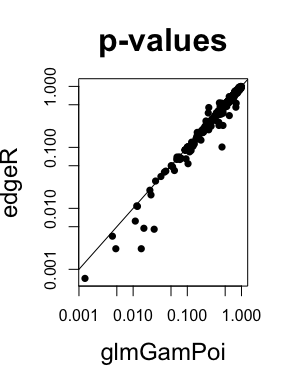
#> 6 ENSG00000151366 0.4037263 0.8776658 0.69733318 1 4431.149 NAThe p-values agree well with the ones that edgeR is calculating. This
is because glmGamPoi uses the same framework of quasi-likelihood ratio
tests that was invented by edgeR and is described in Lund et
al. (2012).
model_matrix <- model.matrix(~ group, data = data.frame(group = group))
edgeR_data <- edgeR::DGEList(pbmcs_subset)
edgeR_data <- edgeR::calcNormFactors(edgeR_data)
edgeR_data <- edgeR::estimateDisp(edgeR_data, design = model_matrix)
edgeR_fit <- edgeR::glmQLFit(edgeR_data, design = model_matrix)
edgeR_test <- edgeR::glmQLFTest(edgeR_fit, coef = 2)
edgeR_res <- edgeR::topTags(edgeR_test, sort.by = "none", n = nrow(pbmcs_subset))Be very careful how you interpret the p-values of a single cell experiment. Cells that come from one individual are not independent replicates. That means that you cannot turn your RNA-seq experiment with 3 treated and 3 control samples into a 3000 vs 3000 experiment through single cell experiment with 1000 cells per sample. The actual unit of replication are still the 3 samples in each condition.
Nonetheless, single cell data is valuable because it allows you to
compare the effect of a treatment on specific cell types. The simplest
way to do such a test is called pseudobulk. This means that the data is
subset to the cells of a specific cell type. Then the counts of cells
from the same sample are combined to form a “pseudobulk” sample. The
test_de() function of glmGamPoi supports this feature directly through
the pseudobulk_by and subset_to parameters:
# say we have cell type labels for each cell and know from which sample they come originally
sample_labels <- rep(paste0("sample_", 1:6), length = ncol(pbmcs_subset))
cell_type_labels <- sample(c("T-cells", "B-cells", "Macrophages"), ncol(pbmcs_subset), replace = TRUE)
test_de(fit, contrast = Group1 - Group2,
pseudobulk_by = sample_labels,
subset_to = cell_type_labels == "T-cells",
n_max = 4, sort_by = pval, decreasing = FALSE)
#> name pval adj_pval f_statistic df1 df2 lfc
#> 110 ENSG00000134539 0.04960240 0.9999771 5.284968 1 8.250891 11.065413
#> 218 ENSG00000158411 0.05579166 0.9999771 4.948362 1 8.250891 -15.112191
#> 300 ENSG00000188243 0.05654532 0.9999771 4.910629 1 8.250891 4.728711
#> 74 ENSG00000217128 0.07338284 0.9999771 4.204576 1 8.250891 13.096497sessionInfo()
#> R version 4.0.0 Patched (2020-05-04 r78358)
#> Platform: x86_64-apple-darwin17.0 (64-bit)
#> Running under: macOS Mojave 10.14.6
#>
#> Matrix products: default
#> BLAS: /Library/Frameworks/R.framework/Versions/4.0/Resources/lib/libRblas.dylib
#> LAPACK: /Library/Frameworks/R.framework/Versions/4.0/Resources/lib/libRlapack.dylib
#>
#> locale:
#> [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
#>
#> attached base packages:
#> [1] parallel stats4 stats graphics grDevices utils datasets
#> [8] methods base
#>
#> other attached packages:
#> [1] TENxPBMCData_1.6.0 HDF5Array_1.16.0
#> [3] rhdf5_2.32.0 SingleCellExperiment_1.10.1
#> [5] DelayedMatrixStats_1.10.0 SummarizedExperiment_1.18.1
#> [7] DelayedArray_0.14.0 matrixStats_0.56.0-9000
#> [9] Biobase_2.48.0 GenomicRanges_1.40.0
#> [11] GenomeInfoDb_1.24.0 IRanges_2.22.1
#> [13] S4Vectors_0.26.0 BiocGenerics_0.34.0
#> [15] glmGamPoi_1.1.7
#>
#> loaded via a namespace (and not attached):
#> [1] bitops_1.0-6 bit64_0.9-7
#> [3] RColorBrewer_1.1-2 httr_1.4.1
#> [5] tools_4.0.0 utf8_1.1.4
#> [7] R6_2.4.1 DBI_1.1.0
#> [9] colorspace_1.4-1 tidyselect_1.1.0
#> [11] DESeq2_1.29.4 bit_1.1-15.2
#> [13] curl_4.3 compiler_4.0.0
#> [15] cli_2.0.2 scales_1.1.0
#> [17] bench_1.1.1 genefilter_1.70.0
#> [19] rappdirs_0.3.1 stringr_1.4.0
#> [21] digest_0.6.25 rmarkdown_2.1
#> [23] XVector_0.28.0 pkgconfig_2.0.3
#> [25] htmltools_0.4.0 dbplyr_1.4.3
#> [27] fastmap_1.0.1 limma_3.44.1
#> [29] rlang_0.4.6 RSQLite_2.2.0
#> [31] shiny_1.4.0.2 generics_0.0.2
#> [33] BiocParallel_1.22.0 dplyr_1.0.0
#> [35] RCurl_1.98-1.2 magrittr_1.5
#> [37] GenomeInfoDbData_1.2.3 Matrix_1.2-18
#> [39] fansi_0.4.1 Rcpp_1.0.4.6
#> [41] munsell_0.5.0 Rhdf5lib_1.10.0
#> [43] lifecycle_0.2.0 stringi_1.4.6
#> [45] yaml_2.2.1 edgeR_3.27.8
#> [47] zlibbioc_1.34.0 BiocFileCache_1.12.0
#> [49] AnnotationHub_2.20.0 grid_4.0.0
#> [51] blob_1.2.1 promises_1.1.0
#> [53] ExperimentHub_1.14.0 crayon_1.3.4
#> [55] lattice_0.20-41 profmem_0.5.0
#> [57] beachmat_2.4.0 splines_4.0.0
#> [59] annotate_1.66.0 locfit_1.5-9.4
#> [61] knitr_1.28 pillar_1.4.4
#> [63] geneplotter_1.66.0 XML_3.99-0.3
#> [65] glue_1.4.0 BiocVersion_3.11.1
#> [67] evaluate_0.14 BiocManager_1.30.10
#> [69] vctrs_0.3.1 httpuv_1.5.2
#> [71] gtable_0.3.0 purrr_0.3.4
#> [73] assertthat_0.2.1 ggplot2_3.3.0
#> [75] xfun_0.13 mime_0.9
#> [77] xtable_1.8-4 later_1.0.0
#> [79] survival_3.1-12 tibble_3.0.1
#> [81] AnnotationDbi_1.50.0 memoise_1.1.0
#> [83] ellipsis_0.3.0 interactiveDisplayBase_1.26.0
#> [85] BiocStyle_2.16.0