This package provides stand-alone version of unet segmentation of fluorescent nuclei based on the code from the Cellprofiler plugin https://github.com/CellProfiler/CellProfiler-plugins/blob/master/classifypixelsunet.py . This is mainly code from the Cellprofiler team (see the Acknowledgements section below).
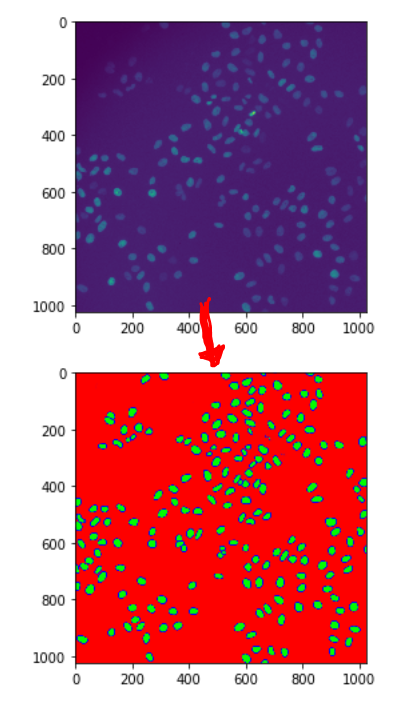
The package comes bundled with a pre-trained Unet to segment fluorescece microsopy images of cell nuclei. The output of the network is a three-channel classification result that provides the probabilities for a pixel to be part of a background, nucleus and boundary pixel. The seperate boundary class is useful for splitting clumped nuclei.
This package was mainly created for personal use, so I can integrate the nuclei segmentation in my own python code without CellProfiler as a dependency. I'm putting it on Github in case others find it useful.
To install use pip (you may have to use pip3 if your default pip refers to python 2.7):
pip install git+https://github.com/VolkerH/unet-nuclei.git
Or clone the repo and install using setup.py:
git clone https://github.com/VolkerH/unet-nuclei.git
python setup.py install
Note that the requirements.txt file does not specify any backends for Keras (see comments in the next section) as there are several options. Therefore you have to install one of the following backend packages using pip or conda and select it as as explained in the following section.
- plaidml-keras (recommended if you have a GPU that is not supported by CUDA, i.e. many laptop GPUs and AMD graphic cards. Also works with CUDA). Don't forget to run
plaidml-setuponce after installingplaidml-keras. - tensorflow-gpu (recommended if you have CUDA.)
- cntk (not sure why one would use this. requires CUDA and Windows)
- tensorflow (CPU-based tensorflow. Too slow to be of practical use for segmenting reasonably sized images)
The unet is implemented using Keras, a high-level python package for describing neural network architectures.
Keras can use different backends for implementing these networks. While tensorflow is the most widespread
backend, there are other options. In particular worth mentioning is the option to use PlaidML. Using PlaidML you can leverage GPU acceleration for many GPUs that do not have CUDA drivers, including many of the GPUs found in laptops. For instructions on how to set up PlaidML as a Keras backend see these installation notes.
Select the backend by setting the environment variable KERAS_BACKEND to either
plaidml.keras.backendfor PlaidMLtensorflowif you have installedtensorflow-gpuortensorflowcntk
You can set the variable either from the calling shell or in your python code using
import os
os.environ["KERAS_BACKEND"] = "plaidml.keras.backend" # or "tensorflow" or "cntk"
Other options to set the backend are described here: https://keras.io/backend/.
See the example Jupyter notebook under ./notebook/test_unet_nuclei.ipynb.
The code is under a BSD-3 License, for the core part see CellProfiler/CellProfiler-plugins#72.
As mentioned in the Overview section, the core functionality of this package is based on code by the Cellprofiler team, specifically based on this publication by Juan Caicendo, Tim Becker, Claire McQuinn et al. They also provided the pre-trained weights for the model. I derived the code from the Cellprofiler Unet plugin but the original code for the publication is here.
The code snippets for determining valid shapes to feed into the unet are taken from Eric Czech's reply to this Gitub issue.
The sample images in the examples directory are taken from the test set of the EMBO High-Troughput screening course 2012 (Beate Neumann and Jutta Bulkescher).
Github issues and pull requests are welcome.
March 2019, Volker Hilsenstein