Visualisation methods and functions for exploring Bayesian Additive Regression Trees (BART). See BART on CRAN, in particular the vignette.
This package is currently under development. Slides from OzViz 2019 workshop on eatmyshorts are available here.
The first occurrence of the famous "eat my shorts" catch phrase was in S01E02 of The Simpsons. In this episode Edna Krabappel also says "visualise it BART" when Bart is having problems with an aptitude test. This clip from the episode summarises how I sometimes feel about visualising BART models.
Below is a walk through of the packages current capabilities, and will be updated as the package develops.
First we fit the BART model on two variables (for simplicity). Note that we are only returning 5 trees from the 1000 post-burnin samples, and only using 10 trees in total.
suppressPackageStartupMessages({
library(eatmyshorts)
library(dplyr)
library(ggplot2)
library(BART)
library(tidytree)
})
Boston <- MASS::Boston
X <- Boston[, c(6, 13)]
y <- Boston$medv
set.seed(99)
bart_model <- wbart(x.train = X,
y.train = y,
nskip = 1000,
ndpost = 1000,
nkeeptreedraws = 5, # keep only 5 mcmc iteration sum-of-trees
ntree = 10, # use only 10 trees in each iteration (default 200)
printevery = 1000
) The current methods for extracting the trees from the BART models are summarise below.
# Current methods for extracting trees:
tidytree_simple <- as_tidytree(bart_model)
tidytree_simple
#> # A tibble: 182 x 5
#> # Groups: iter, tree_id [50]
#> iter tree_id node parent label
#> <int> <int> <int> <int> <chr>
#> 1 1 1 1 NA rm < 6.3
#> 2 1 1 2 1 -1.06
#> 3 1 1 3 1 lstat < 11.78
#> 4 1 1 6 3 -0.3
#> 5 1 1 7 3 -4.07
#> 6 1 2 1 NA rm < 7.64
#> 7 1 2 2 1 4.75
#> 8 1 2 3 1 8.7
#> 9 1 3 1 NA lstat < 16.44
#> 10 1 3 2 1 -0.35
#> # … with 172 more rows
tidytree_detailed <- as_tidytree(
bart_model,
extra_cols = c("var", "cut", "leaf_value", "is_leaf")
)
tidytree_detailed
#> # A tibble: 182 x 9
#> # Groups: iter, tree_id [50]
#> iter tree_id node parent label var cut leaf_value is_leaf
#> <int> <int> <int> <int> <chr> <chr> <dbl> <dbl> <lgl>
#> 1 1 1 1 NA rm < 6.3 rm 6.30 NA FALSE
#> 2 1 1 2 1 -1.06 <NA> NA -1.06 TRUE
#> 3 1 1 3 1 lstat < 11.78 lstat 11.8 NA FALSE
#> 4 1 1 6 3 -0.3 <NA> NA -0.300 TRUE
#> 5 1 1 7 3 -4.07 <NA> NA -4.07 TRUE
#> 6 1 2 1 NA rm < 7.64 rm 7.64 NA FALSE
#> 7 1 2 2 1 4.75 <NA> NA 4.75 TRUE
#> 8 1 2 3 1 8.7 <NA> NA 8.70 TRUE
#> 9 1 3 1 NA lstat < 16.44 lstat 16.4 NA FALSE
#> 10 1 3 2 1 -0.35 <NA> NA -0.351 TRUE
#> # … with 172 more rows
tidygraph_detailed <- as_tidygraph_list(
bart_model,
extra_cols = c("var", "cut", "leaf_value", "is_leaf"))
tidygraph_detailed[1:2]
#> [[1]]
#> # A tbl_graph: 5 nodes and 4 edges
#> #
#> # A rooted tree
#> #
#> # Node Data: 5 x 8 (active)
#> node label var cut leaf_value is_leaf iter tree_id
#> <int> <chr> <chr> <dbl> <dbl> <lgl> <int> <int>
#> 1 1 rm < 6.3 rm 6.30 NA FALSE 1 1
#> 2 2 -1.06 <NA> NA -1.06 TRUE 1 1
#> 3 3 lstat < 11.78 lstat 11.8 NA FALSE 1 1
#> 4 4 -0.3 <NA> NA -0.300 TRUE 1 1
#> 5 5 -4.07 <NA> NA -4.07 TRUE 1 1
#> #
#> # Edge Data: 4 x 2
#> from to
#> <int> <int>
#> 1 1 2
#> 2 1 3
#> 3 3 4
#> # … with 1 more row
#>
#> [[2]]
#> # A tbl_graph: 3 nodes and 2 edges
#> #
#> # A rooted tree
#> #
#> # Node Data: 3 x 8 (active)
#> node label var cut leaf_value is_leaf iter tree_id
#> <int> <chr> <chr> <dbl> <dbl> <lgl> <int> <int>
#> 1 1 rm < 7.64 rm 7.64 NA FALSE 1 2
#> 2 2 4.75 <NA> NA 4.75 TRUE 1 2
#> 3 3 8.7 <NA> NA 8.70 TRUE 1 2
#> #
#> # Edge Data: 2 x 2
#> from to
#> <int> <int>
#> 1 1 2
#> 2 1 3The tidytree method returns a tbl_tree (special tibble) grouped by MCMC iteration and tree number, whilst the tidygraph method returns a list of tbl_graph which inherit from the igraph structure.
By using the ggraph and tidygraph package together, we can do some simple plotting.
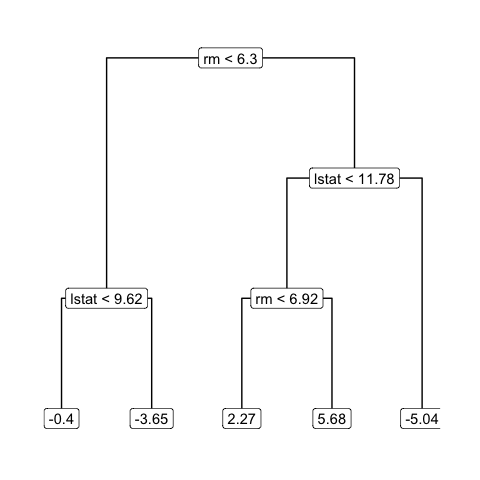
library(ggraph)
ggraph(tidygraph_detailed[[31]], 'dendrogram') +
geom_edge_elbow() +
geom_node_label(aes(label = label)) +
theme_graph()This only plots 1 tree from 1 iteration of the BART MCMC. How do we extend it?