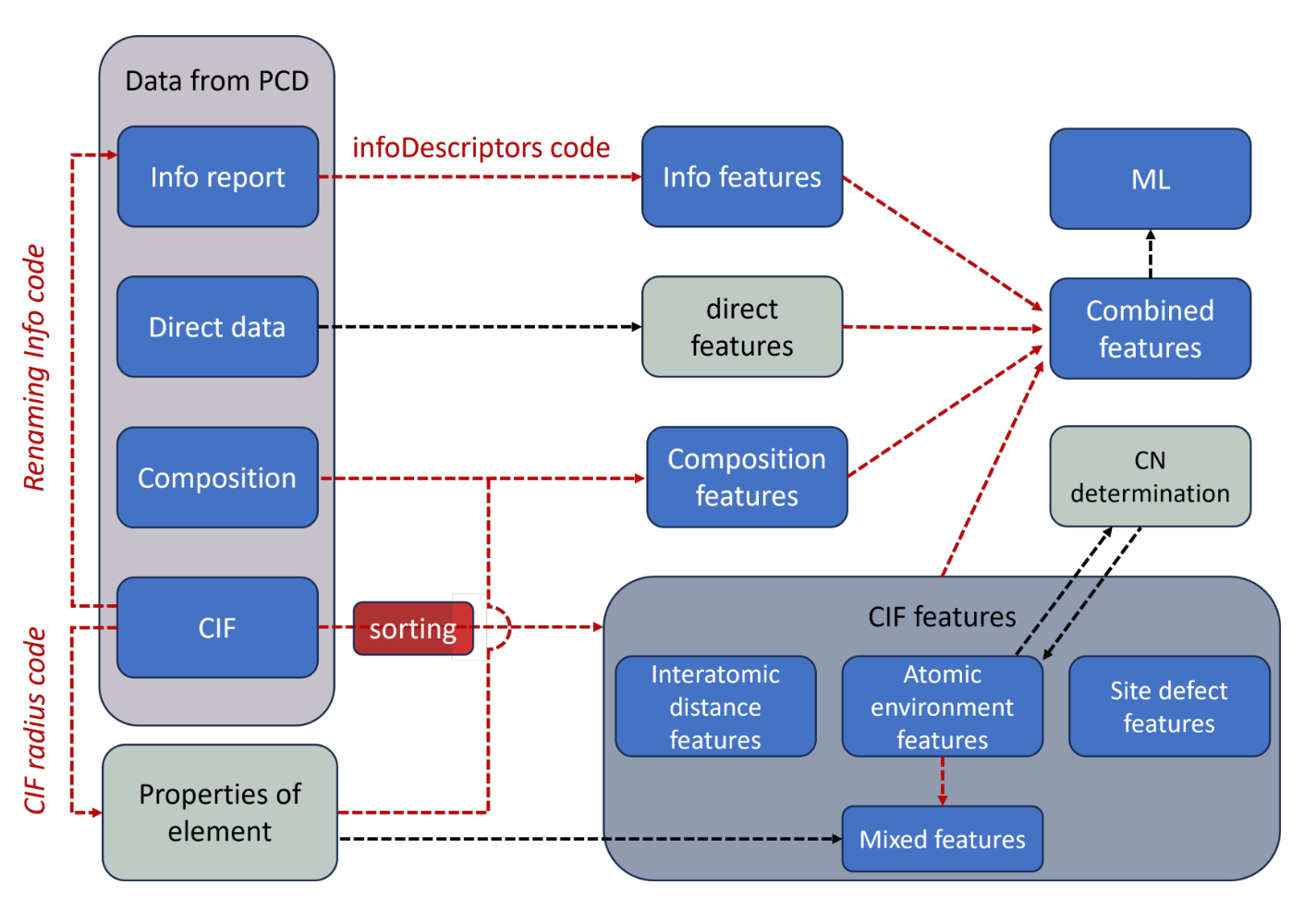
Structure Analysis/Featurizer (SAF) is a Python script designed to process CIF (Crystallographic Information File) files and extract geometric features. These features include interatomic distances, information on atomic environments, and coordination numbers. The script can sequentially process more than 10,000 .cif files and generate millions of data points used as ML input data for predicting crystal structures and properties.
The script was originally developed to determine the coordination number for each crystallographic site on complex structures [1]. Then, we further included an interactive functionality for experimentalists and data scientists to generate structural features from .cif file. These features were engineered to be used as input data for ML models to predict crystal structures and their properties. We are currently exploring this option in Summer 2024.
Start with the script via
python main.pyOnce the script is started, the user needs to only confirm (1) whether to skip cif files based on the number of atoms in the supercell generated and (2) choose the folder.
Welcome to CIF Featurizer! This script processes Crystallographic Information
File (CIF) files to extract various features such as interatomic distances,
atomic environment information, and coordination numbers. It supports binary
and ternary compounds.
Q1. Do you want to skip any CIF files based on the number of unique in the supercell?
(Default: N) [y/N]: N
Available folders containing CIF files:
1. 20240226_huge_file_test
2. 20240402_ThSb
3. 20240303_ternary_binary_test
4. 20240402_URhIn
Enter the number corresponding to the folder containing .cif files: 3
After running the script using python main.py and selecting the folder contianing .cif files, .csv files are generated. For binary compounds, feature_binary.csv with 124 features is generated. For ternary compounds, feature_ternary.csv with 165 unique features is generated. For all types of compounds, feature_universal.csv is generated.
The coordination numbers and their geometric values are determined using four unique coordination determination methods, as detailed in the manuscript (DOI). Below is an example from a site. Once the code becomes more matured, we will further make a better documentation here.
| # | CN method | Central Atom | CN | R | M | X | Polyhedron volume | Dist from atom to center of mass | Edges | Faces |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | Shortest dist | Er1 | 13 | 5 | 2 | 6 | 89.293 | 0.095 | 33 | 22 |
| 2 | CIF radius sum | Er1 | 13 | 5 | 2 | 6 | 89.293 | 0.095 | 33 | 22 |
| 3 | CIF radius refined sum | Er1 | 13 | 5 | 2 | 6 | 89.293 | 0.095 | 33 | 22 |
| 4 | Pualing radius sum | Er1 | 13 | 5 | 2 | 6 | 89.293 | 0.095 | 33 | 22 |
Before running the script, make sure you have the following dependencies installed:
pip install click gemmi matplotlib numpy openpyxl pandas pytest scipy sympy
cd cif-featurizer
python main.pyThe recommended way for installation is Conda
git clone https://github.com/bobleesj/structure-analyzer-featurizer.git
cd structure-analyzer-featurizer
conda create -n cif python=3.12
conda activate cif
pip install -r requirements.txt
python main.pyIf you are new to Conda (Python package manager), you may refer to Intro to Python package manager for beginners (Ft. Conda with Cheatsheet.
SAF provides built-in preprocessing in this workflow to manually sort out .cif files with incorrect formatting. You may further use CIF Cleaner to formats .cif files and sorts them based on tags, supercell size, and minimum distance to save time.
- Anton Oliynyk - CUNY Hunger College
- Arnab Dutta - IIT Kharagpur
- Nikhil Kumar Barua - University of Waterloo
- Nishant Yadav - IIT Kharagpur
- Sangjoon Bob Lee - Columbia University
- Siddha Sankalpa Sethi - IIT Kharagpur
Here is a list of publications that have used this code for analysis:
[1] Y. Tyvanchuk, V. Babizhetskyy, S. Baran, A. Szytula, V. Smetana, S. Lee, A. O. Oliynyk, A. Mudring, The crystal and electronic structure of RE23Co6.7In20.3 (RE = Gd–Tm, Lu): A new structure type based on intergrowth of AlB2- and CsCl-type related slabs. Journal of Alloys and Compounds. 976, 173241 (2024). doi.org/10.1016/j.jallcom.2023.173241