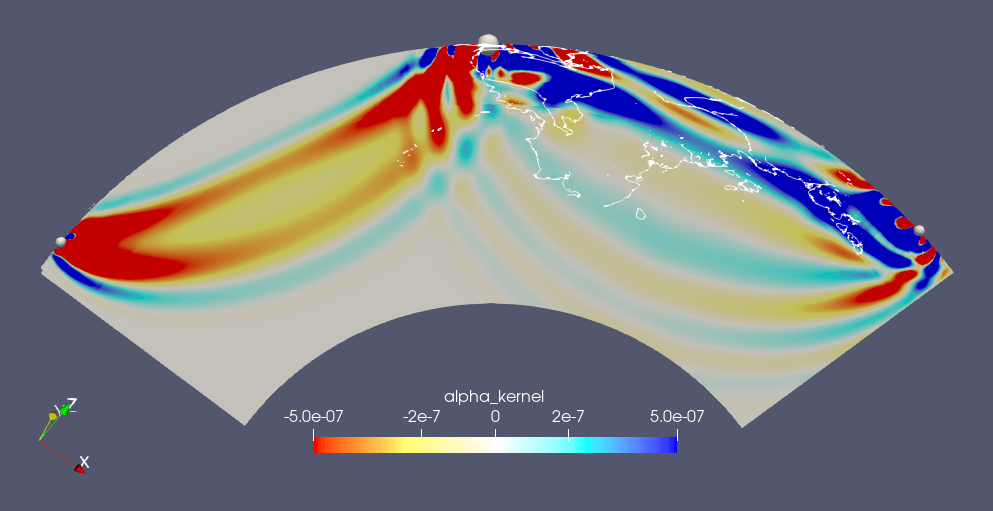
This repository contains the jupyter notebook and data for the 2024 SCOPED workshop. Below is a brief description of the files in this repository:
data_processing_and_kernel_comp.ipynb: A jupyter notebook that demonstrates how to process the data and run a kernel computation.data_processing_and_kernel_comp_run_on_local.ipynb: The same notebook as the above, but it is designed to run on local machines with docker.data: the data for the demo, which will be downloaded in the notebook.quakeml: the QuakeML files, which will be created after running the notebook.simulation: a directory where we run forward/adjoint simulations with Specfem3D_globe. The essential files will be created here after running the notebook.img: images used in the notebook and README.md.shakemov_syn: the synthetic waveform data downloaded from the ShakeMovie website.finite_fault: the CMTSOLUTION files for the finite fault model.job.jupyter: a job script for running the jupyter notebook on Frontera.job.dcv: a job script for running the visualization job on Frontera.create_slice.py: paraview python script for creating slices.AVS_boundaries_elliptical.inp: an AVS input file for plotting the coastlies.paraview_red_to_blue_colormap.json: a paraview colormap file for plotting kernel.plot_kernel_slices_frontera.pvsm: a paraview state file for plotting kernel slices.
Some parts of this example are designed to run on the Frontera supercomputer at TACC. If you don't have access to Frontera, but have a docker installed on your local machine, you can run the notebook including forward and adjoint simulation with Specfem3D_globe on your local machine. Please follow the section here section. If you don't have access to Frontera or no docker installation on your machine, you can still run the notebook on your local machine to do a data processing and calculation adjoint source. For running this on your local machine, please follow the section here section.
This example is designed to run on the Frontera supercomputer at TACC. To log in to Frontera, you need to have an account at TACC and authentication setup. If you don't have an account, please follow the instruction to setup it here.
- go to SCRATCH directory:
cd $SCRATCH- Clone this repository:
git clone https://github.com/mnagaso/workshop_scoped_2024_specfem3d_globe.git- Submit the job:
cd workshop_scoped_2024_specfem3d_globe
sbatch job.jupyter- Check the job status:
squeue -u $USER
JOBID PARTITION NAME USER ST TIME NODES NODELIST(REASON)
6247768 development tap_jupy mnagaso PD 0:00 1 (None)- You will find the starting of the job by becoming
RfromPDfor example:
squeue -u $USER
JOBID PARTITION NAME USER ST TIME NODES NODELIST(REASON)
6247768 development tap_jupy mnagaso R 0:00 1 c201-022It takes about 1 mintue or so to finishing a initial setup. Then, you can access the jupyter notebook server by openning the link indicated at the last of the jupyter.out file.
This link is something like:
tail jupyter.out
TACC: created reverse ports on Frontera logins
TACC: Your jupyter notebook server is now running at https://frontera.tacc.utexas.edu:60188/?token=ee7153b2ec3569dabea24b66de63247efed8cf2e8f203036cc2f490c58321fc7
Stop current job for jupyter notebook by running the command below on the terminal:
scancel -u $USERThen, start a new job for the visualization by running the command below on the terminal:
sbatch ./job.dcvAfter the job is started, you will have the url for opening the visualization job environment, at the end of the output file dcvserver.out, e.g.
TACC: Your DCV session is now running!
TACC: To connect to your DCV session, please point a modern web browser to:
TACC: https://frontera.tacc.utexas.edu:60036
You can load the paraview module and the state file for plotting the kernel slices by running the command below on the terminal:
./run_visualization.shThis example may be run on the computers other than Frontera. If you have a docker installed on your machine with sufficient RAM and disk space, you can run the notebook including forward and adjoint simulation with Specfem3D_globe on your local machine. Please follow the instruction below.
Please note that the default setup for this setup takes more than 5 hours with 4 MPI processes, so you cannot finish the calculation within the workshop time frame, uneless you have a powerful machine and increase the number of MPI processes.
(x86_64 architecture is required for running the docker image, as the base image is not supported on ARM architecture.)
docker pull ghcr.io/mnagaso/specfem3d_globe:centos7_mpiYou may verify the image is downloaded by running the command below:
$ docker images
REPOSITORY TAG IMAGE ID CREATED SIZE
ghcr.io/mnagaso/specfem3d_globe centos7_mpi 6cab8b5522e7 3 days ago 5.55GBpip install --user obspy cartopy jupyterlabpip uninstall -y urllib3
pip install --user 'urllib3<2.0'jupyter-lab
or
jupyter-notebook
or
jupyter lab
or
jupyter notebookOpen data_processing_and_kernel_comp_run_on_local.ipynb on the opened jupyter GUI.
If you don't have access to Frontera, you can still run the notebook on your local machine. Here is a brief instruction on how to run the notebook on your local machine:
pip install --user obspy cartopy jupyterlabpip uninstall -y urllib3
pip install --user 'urllib3<2.0'jupyter-lab
or
jupyter-notebook
or
jupyter lab
or
jupyter notebookPlease modify the line in the data_process_and_kernel_comp.ipynb (7th cell) for loading pre-calculated synthetic data as below (to load the precalculated synthetics):
# read and plot the raw data
...
# load the synthetic waveform data with 3D model (with SPECFEM3D_GLOBE)
# use the result from the forward simulation
#st_syn = obspy.read("./simulation/OUTPUT_FILES/*sac") # in sac format
# use the precalculated synthetics with 3D model
st_syn = obspy.read("./data/C202401010710A/waveforms_syn/*sac") # in sac format
...